The Thing Metabolome Repository family (XMRs): comparable untargeted metabolome databases for analyzing sample-specific unknown metabolites
Press release
The Thing Metabolome Repository family (XMRs): comparable untargeted metabolome databases for analyzing sample-specific unknown metabolites
Nozomu Sakurai*†, Shinichi Yamazaki†, Kunihiro Suda, Ai Hosoki, Nayumi Akimoto, Haruya Takahashi, Daisuke Shibata and Yuichi Aoki*
* Corresponding authors† First authors
Nucleic Acids Research (Database Issue) 2022 November 24 DOI:10.1093/nar/gkac1058
![]() Press release (In Japanese only)
Press release (In Japanese only)
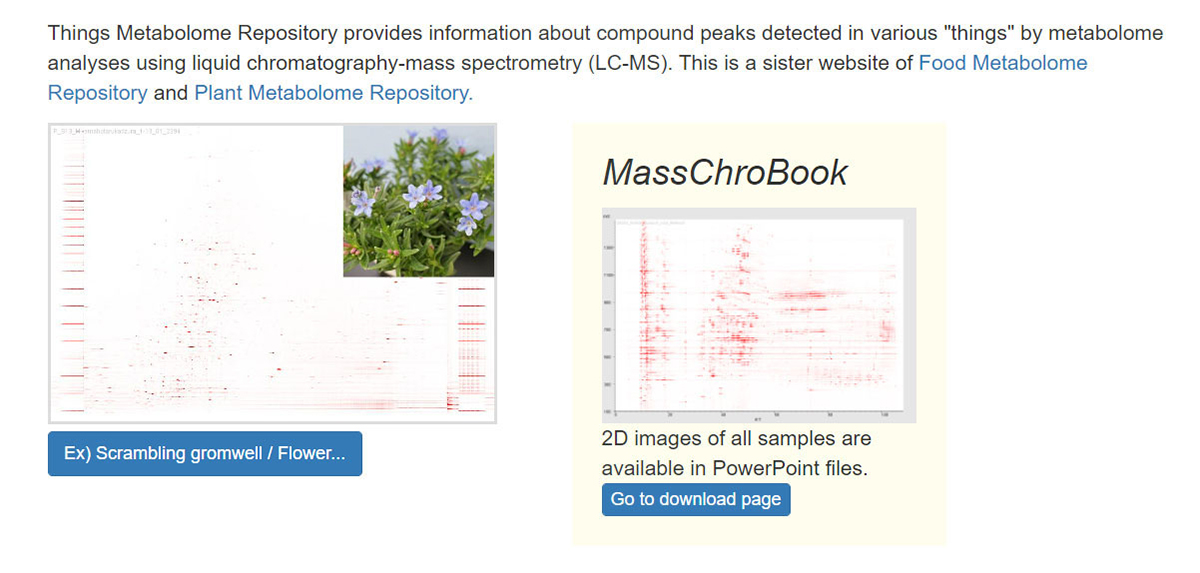
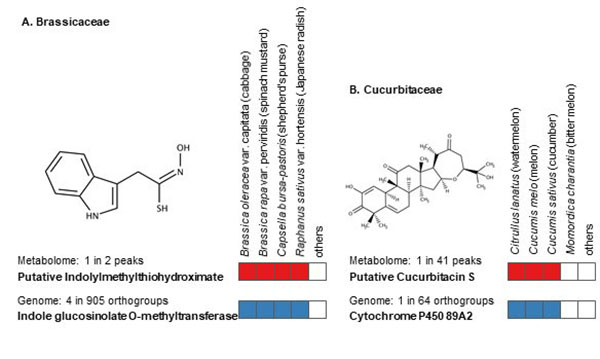
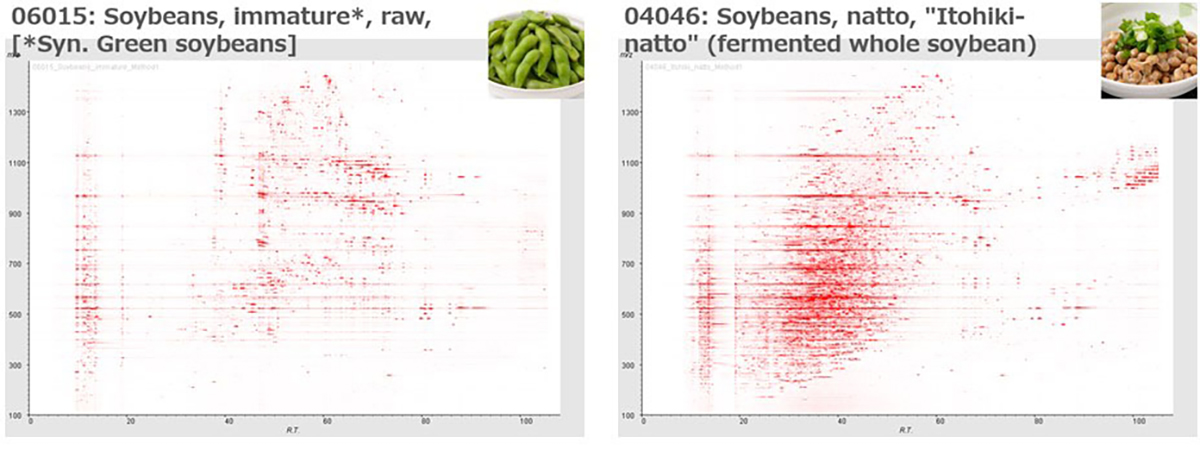
The identification of unknown chemicals has emerged as a significant issue in untargeted metabolome analysis owing to the limited availability of purified standards for identification; this is a major bottleneck for the accumulation of reusable metabolome data in systems biology. Public resources for discovering and prioritizing the unknowns that should be subject to practical identification, as well as further detailed study of spending costs and the risks of misprediction, are lacking. As such a resource, we released databases, Food-, Plant-, and Thing- Metabolome Repository (http://metabolites.in/foods, http://metabolites.in/plants, and http://metabolites.in/things, referred to as XMRs) in which the sample-specific localization of unknowns detected by liquid chromatography–mass spectrometry in a wide variety of samples can be examined, helping to discover and prioritize the unknowns. A set of application programming interfaces for the XMRs facilitates the use of metabolome data for large-scale analysis and data mining. Several applications of XMRs, including integrated metabolome and genome analyses, are presented. Expanding the concept of XMRs will accelerate the identification of unknowns and increase the discovery of new knowledge.
Source: N. Sakurai et al., DOI: 10.1093/nar/gkac1058
▶ FoodMR
▶ PlantMR
▶ ThingMR