Liquid-like chromatin in the cell and the cohesin activity
Maeshima Group / Genome Dynamics Laboratory
Liquid-like chromatin in the cell: What can we learn from imaging and computational modeling?
Yuji Itoh, Esmae J. Woods, Katsuhiko Minami, Kazuhiro Maeshima, and Rosana Collepardo-Guevara
Current Opinion in Structural Biology 71, 123-135 (2021) DOI:10.1016/j.sbi.2021.06.004
The loopy world of cohesin.
Kazuhiro Maeshima and Shiori Iida
eLife 10, e71585 (2021) DOI:10.7554/eLife.71585
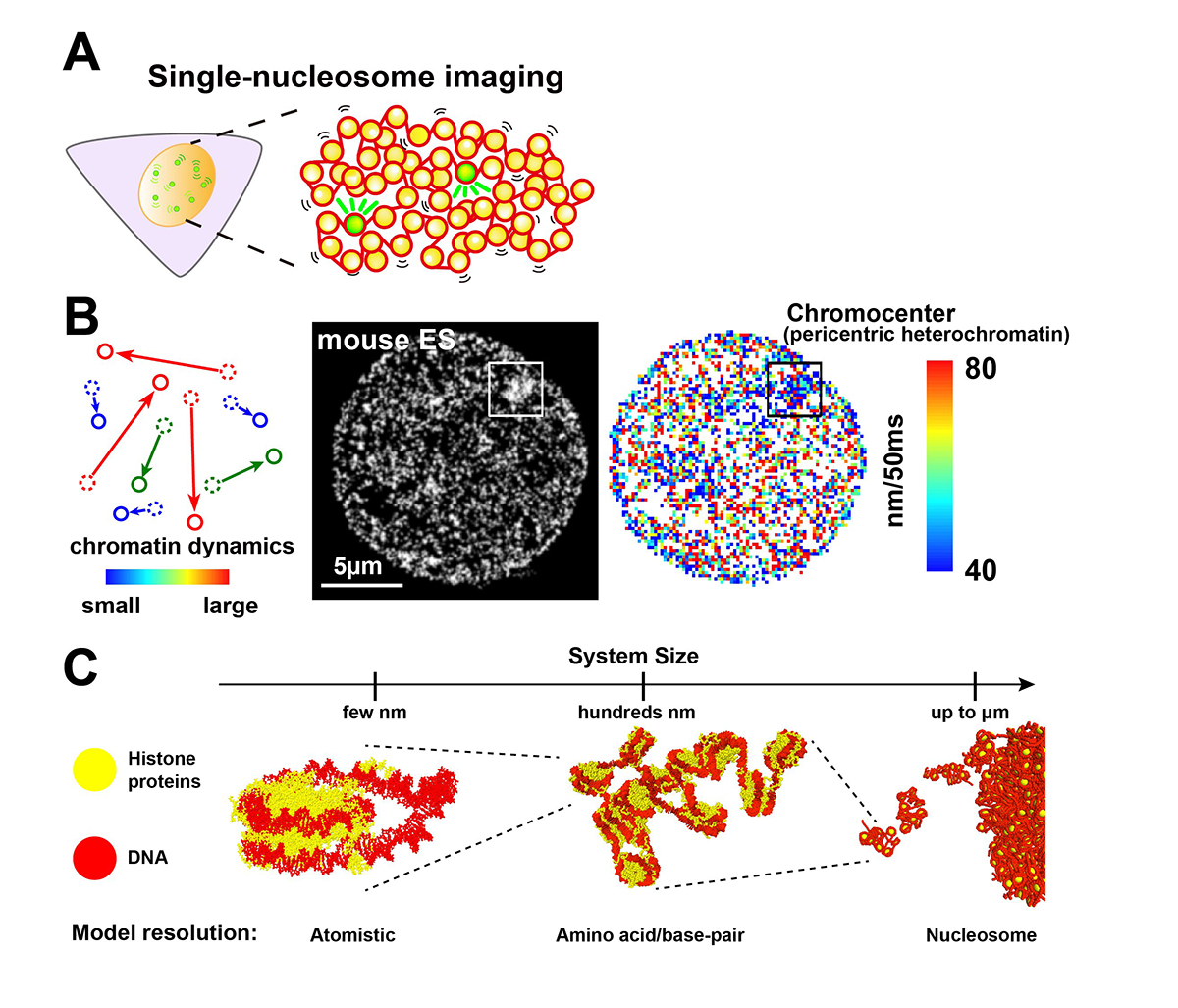
Chromatin in eukaryotic cells is a negatively charged long polymer consisting of DNA, histones, and various associated proteins. With its highly charged and heterogeneous nature, chromatin structure varies greatly depending on various factors (e.g., chemical modifications and protein enrichment) and the surrounding environment (e.g., cations): From a 10-nm fiber, a folded 30-nm fiber, to chromatin condensates/droplets. Recent advanced imaging such as single-nucleosome imaging (Figure 1A) has observed that chromatin exhibits a dynamic liquid-like behavior and undergoes structural variations within the cell (Figure 1B). Current computational modeling has made it possible to reconstruct the liquid-like chromatin in the cell by dealing with a number of nucleosomes on multi-scale levels, and has become a powerful technique to inspect the molecular mechanisms giving rise to the observed behavior, which imaging methods cannot do on their own (Figure 1C). Based on new findings from both imaging and modeling studies, we discuss the dynamic aspect of chromatin in living cells and its functional relevance.
This work was supported by JSPS grant (19K23735, 20J00572, 20H05936, 21H02453), the Takeda Science Foundation, the Uehara Memorial Foundation, NIG Postdoctoral Fellowship, JSPS Postdoctoral Fellowship (PD).
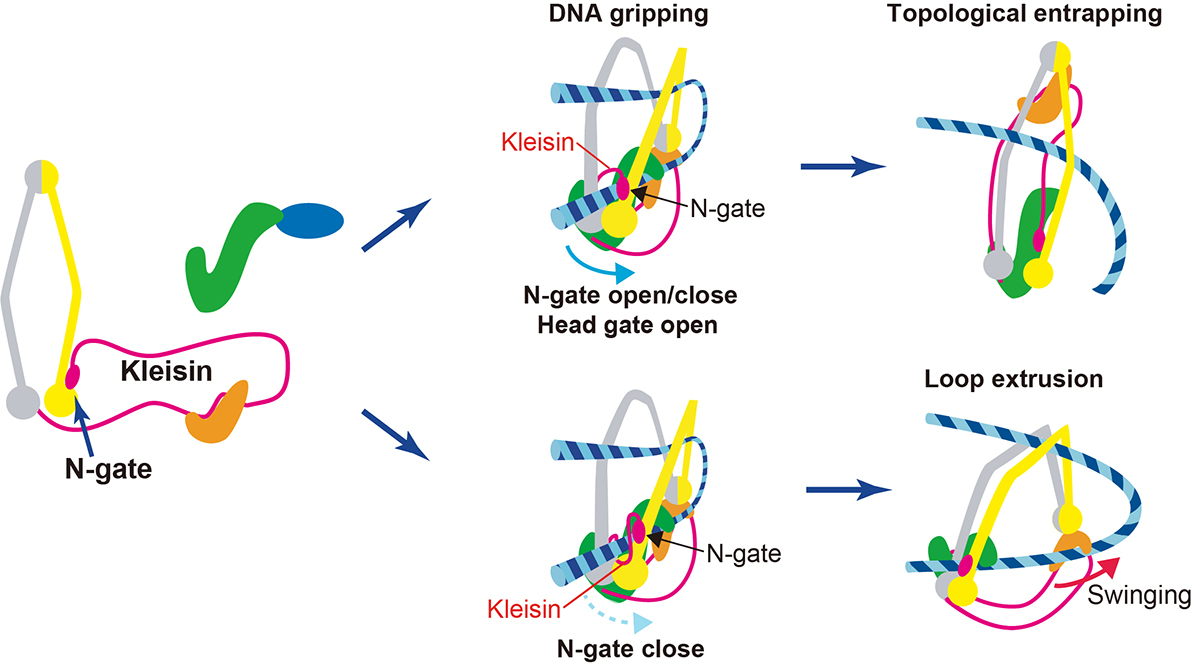
Also, on July 26th, a professor at Genome Dynamics Laboratory Kazuhiro Maeshima and SOKENDAI Ph.D. student Shiori Iida published an Insight paper in eLife. Chromatin higher-order structures, such as chromatin loop domains, are critical for chromatin to perform various functions in the cell. The formation of these chromatin loops is thought to be mediated by a ring-shaped molecular complex, cohesin (Figure 2, left). Currently, the mechanism of chromatin loop formation is a hot topic in cell biology, and a model called loop extrusion, in which cohesin pushes DNA out of the ring, has been attracting much attention. In fact, cohesin has been shown to extrude naked DNA in vitro. Recently, Dr. Frank Uhlmann and his colleagues have shown how cohesin excludes DNA loops in vitro (Figure 2, bottom)(Higashi et al., “A Brownian ratchet model for DNA loop extrusion by the cohesin complex”. eLife, 2021 DOI:10.7554/eLife.67530). Based on the Higashi et al. paper, Maeshima and Iida discussed the intracellular behavior of cohesin and whether the loop extrusion reported really occurs in the cell.