無脊椎動物遺伝研究室
齋藤研究室
エピトランスクリプトームの分子機構と生理機能解明
教員

Research Summary
RNAに見出される多様なRNA修飾は、核酸に新たな化学的性質を付加し様々な分子との機能的連携を制御することで新たな階層のエピトランスクリプトミックコードを形成する。我々は、モデル動物ショウジョウバエ、哺乳類培養細胞を駆使し、時空間の様々なレベルでRNA修飾の分子機構、ダイナミクス、生理機能、を明らかにすることを目指している。
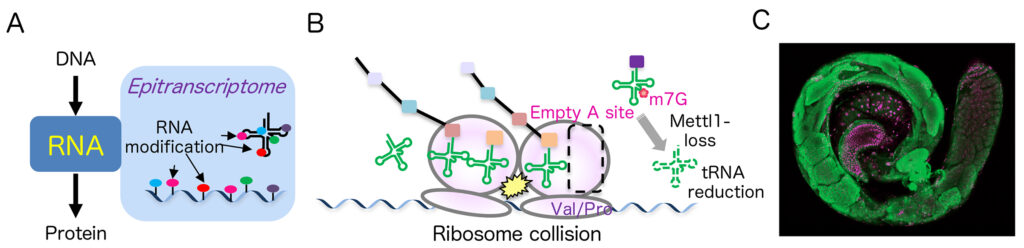
現在、170種以上のRNA修飾が同定され、研究が進められています。
(B)私たちはメチル基転移酵素の一つであるMettl1がtRNAのm7Gメチル化とその量の制御に重要であることを明らかにしました。
(C)Mettl1ノックアウトハエ精巣の蛍光顕微鏡像。伸長精子細胞形成が異常となり、次世代を残せないことを発見しました(Nature Communications 2024)。
出版物
- Kaneko S, Miyoshi K, Tomuro K, Terauchi M, Tanaka R, Kondo S, Tani N, Ishiguro KI, Toyoda A, Kamikouchi A, Noguchi H, Iwasaki S, and Saito K* (2024). Mettl1-dependent m(7)G tRNA modification is essential for maintaining spermatogenesis and fertility in Drosophila melanogaster. Nat Commun 15, 8147. 10.1038/s41467-024-52389-0.
- Yamamoto-Matsuda H, Miyoshi K, Moritoh M, Yoshitane H, Fukada Y, Saito K, Yamanaka S, Siomi MC. Lint-O cooperates with L(3)mbt in target gene suppression to maintain homeostasis in fly ovary and brain. EMBO Rep. 2022 Oct 6;23(10):e53813.
- Utsuno Y, Hamada K, Hamanaka K, Miyoshi K, Tsuchimoto K, Sunada S, Itai T, Sakamoto M, Tsuchida N, Uchiyama Y, Koshimizu E, Fujita A, Miyatake S, Misawa K, Mizuguchi T, Kato Y, Saito K, Ogata K, Matsumoto N. Novel missense variants cause intermediate phenotypes in the phenotypic spectrum of SLC5A6-related disorders. J Hum Genet. 2024 Feb;69(2):69-77.
- Takeuchi C, Yokoshi M, Kondo S, Shibuya A, Saito K, Fukaya T, Siomi H, Iwasaki YW. Mod(mdg4) variants repress telomeric retrotransposon HeT-A by blocking subtelomeric enhancers. Nucleic Acids Res. 2022 Nov 11;50(20):11580-11599.
