微生物機能研究室
仁木研究室
モデル単細胞を使った細胞分裂の遺伝制御メカニズム
教員


Research Summary
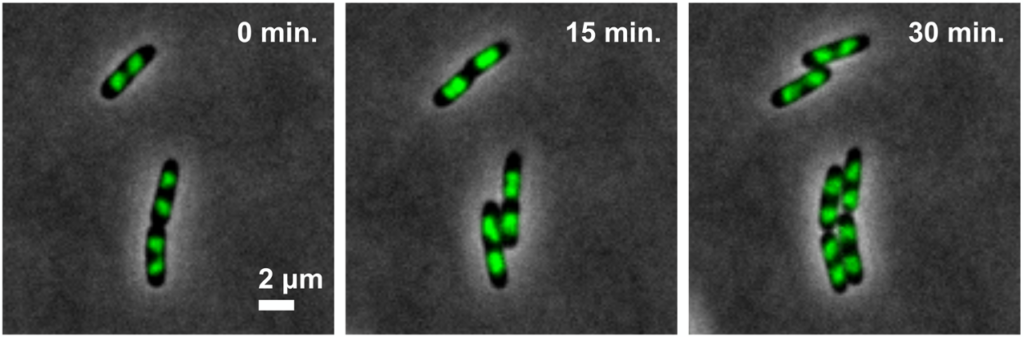
大腸菌や酵母は、細胞増殖の基本メカニズムを解明する上で極めて有効なモデル生物です。原核細胞と真核細胞を研究材料に、染色体DNAの折れたたみを司るコンデンシンの機能、光や温度に対する細胞応答、細胞の形が決まる仕組み、タンパク質が合成される仕組み等の研究を進めています。遺伝学的もしくは細胞生物学的手法を用いて、細胞内で起こる現象を観察しています。特に、酵母と菌糸という2つの生活環を持つジャポニカス分裂酵母は環境刺激に対する細胞応答のモデル細胞として適しています。またDNA組換え技術の宿主として、より優れた大腸菌の開発も担っています。
出版物
- Fujiwara K, Tsuji N, Yoshida M, Takada H, Chiba S. Patchy and widespread distribution of bacterial translation arrest peptides associated with the protein localization machinery. Nat Commun. 2024 Apr 2;15(1):2711.
- Yano K, Noguchi H, Niki H. Profiling a single-stranded DNA region within an rDNA segment that affects the loading of bacterial condensin. iScience. 2022 Nov 4;25(12):105504.
- Seike T, Sakata N, Shimoda C, Niki H, Furusawa C. The sixth transmembrane region of a pheromone G-protein coupled receptor, Map3, is implicated in discrimination of closely related pheromones in Schizosaccharomyces pombe. Genetics. 2021 Dec 10;219(4):iyab150.
- Nakai R, Wakana I, Niki H. Internal microbial zonation during the massive growth of marimo, a lake ball of Aegagropila linnaei in Lake Akan. iScience. 2021 Jun 12;24(7):102720.
