大量遺伝情報研究室
中村研究室
種々の生物を対象とした塩基配列解析とデータベースの高度化
教員

Research Summary
次世代高速シークエンサの技術革新により得られる大量の塩基配列データを活用するためには、高品質な塩基配列の解析と使いやすいデータベースの提供が不可欠です。また信頼性の高い自動化解析システムの開発も重要です。
中村研究室は高度なゲノム情報解析とそのデータベース化や、アノテーションの質の向上に取り組んでいます。日本DNAデータバンク(DDBJ)の業務担当研究室として、原核生物の自動解析システムDFASTを開発し高速かつ正確なデータ解析を支援しています。また、苔類ゼニゴケMarchantia polymorpha、イエネコFelis catus のような進化研究上、産業上あるいは医療上重要な生物種の高精度な解析を実施しています。
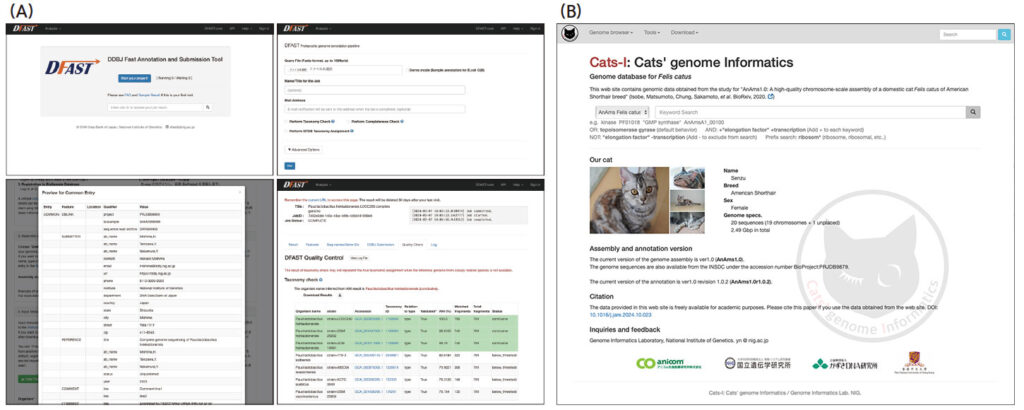
(B)Cats-I:Cats’ genome informaticsイエネコゲノム情報データベース
出版物
- Elmanzalawi M, Fujisawa T, Mori H, Nakamura Y, Tanizawa Y. DFAST_QC: quality assessment and taxonomic identification tool for prokaryotic Genomes. BMC Bioinformatics. 2025 Jan 7;26(1):3.
- Matsumoto Y, Yik-Lok Chung C, Isobe S, Sakamoto M, Lin X, Chan TF, Hirakawa H, Ishihara G, Lam HM, Nakayama S, Sasamoto S, Tanizawa Y, Watanabe A,Watanabe K, Yagura M, Niimura Y, Nakamura Y. Chromosome-scale assembly with improved annotation provides insights into breed-wide genomic structure and diversity in domestic cats. J Adv Res. 2024 Oct 28:S2090-1232(24)00478-8.
- Mochizuki T, Sakamoto M, Tanizawa Y, Nakayama T, Tanifuji G, Kamikawa R,Nakamura Y. A practical assembly guideline for genomes with various levels of heterozygosity. Brief Bioinform. 2023 Sep 22;24(6):bbad337.
- Kawamura S, Romani F, Yagura M, Mochizuki T, Sakamoto M, Yamaoka S,Nishihama R, Nakamura Y, Yamato KT, Bowman JL, Kohchi T, Tanizawa Y. MarpolBase Expression: A Web-Based, Comprehensive Platform for Visualization and Analysis of Transcriptomes in the Liverwort Marchantia polymorpha. Plant Cell Physiol. 2022 Nov 22;63(11):1745-1755.
