分子生命史研究室
工樂研究室
生物多様性を導いた動物進化の分子基盤に迫る
教員



Research Summary
本研究室では、ゲノムDNA配列情報を分子系統学的観点から解析するとともに、ゲノムワイドな分子情報プロファイリングがもたらす細胞レベルの現象の知見を統合し、複雑な生命の成り立ちを理解するための研究を進めています。脊椎動物を中心とした、生物学的に際立つ特徴を持つ希少生物を含む多様な生物を対象としています。主要なテーマは以下の3つに分けられます。
- DNA配列の種間比較によるゲノム構成の進化的変遷の解明
- 細胞レベルの現象の理解に基づくゲノム進化機構の解明
- ゲノムワイドな情報の取得および利用のための方法の高度化
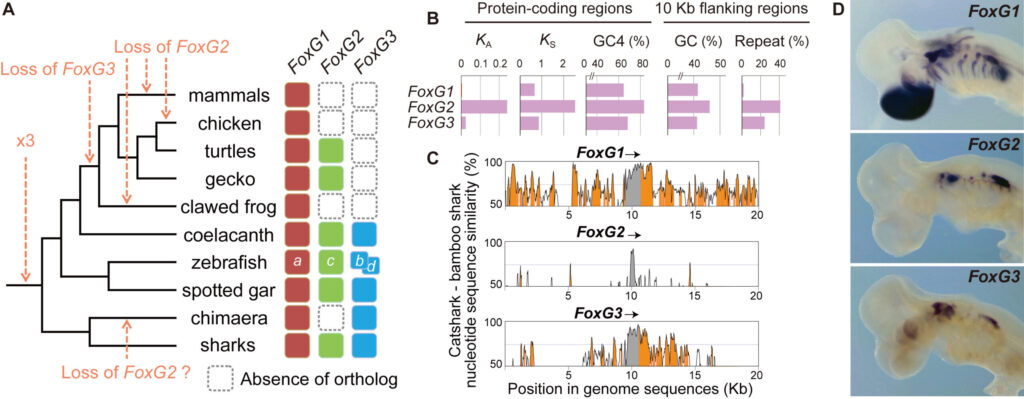
(A)、塩基組成や周辺の反復配列頻度(B)、遺伝子配列と周辺のゲノム配列の進化速度(BとC)、胚発生期の発現部位(D、トラザメ胚)等が異なる。こういった性質の連関や、他の遺伝子群も含めたゲノム全体の情報発現が調和するしくみについては謎がまだ多い。分子進化と表現型進化の橋渡しを目指す当研究室では、こういった謎の解明に挑むべく、特定の現象や遺伝子に絞らず、様々な動物を視野に入れて研究を進めている。
出版物
- Kawaguchi YW, Matsumoto R, Kuraku S, Improved genome assembly of whale shark, the world’s biggest fish: revealing intragenomic heterogeneity in molecular evolution, GigaScience, 2026, giag014,
https://doi.org/10.1093/gigascience/giag014 - Kuraku S. Vertebrate Hox clusters as transposon repellents against the genomic accordion model, Zool. Sci. 43: 35-43.
https://doi.org/10.2108/zs250058 - Niwa T, Uno Y, Ohishi Y, Kadota M, Aburatani N, Kiyatake I, Katooka D, Yorozu M, Tsuzuki N, Toyoda A, Takagi W, Nakamura M, Kuraku S. Sharks and rays have the oldest vertebrate sex chromosome with unique sex determination mechanisms, Proc. Natl. Acad. Sci. U.S.A. 2025, 122:e251367612
https://doi.org/10.1073/pnas.2513676122 - Hara Y, Kuraku S. Intragenomic mutational heterogeneity: structural and functional insights from gene evolution. Trends Genet. 2025, May; 5:S0168-9525(25)00075-7.
https://doi.org/10.1016/j.tig.2025.03.007
