マウス開発研究室
小出研究室
野生由来マウスを用いた行動遺伝学
教員

Research Summary
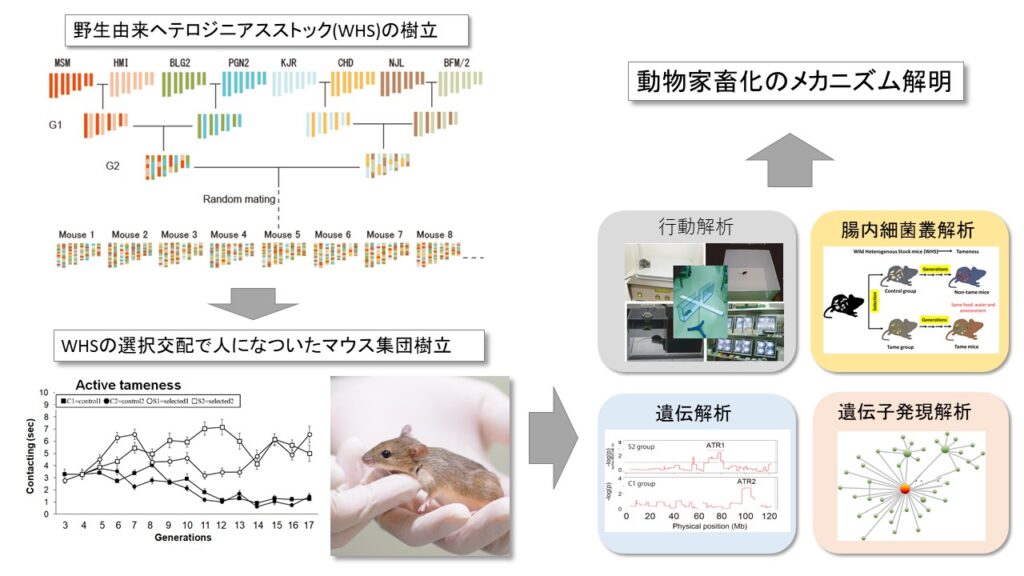
生物の個体差をもたらす遺伝的機構の多くは未だ解明されていません。私たちは、世界各地で捕獲されたマウスをもとに樹立された野生由来系統を用い、様々な行動の多様性を生み出すメカニズムの解明に取り組んでいます。野生由来の近交系統は、特徴的な行動を示し、顕著な系統差を示すことから、行動遺伝学研究に有用です。加えて、ゲノム編集技術を用いた効率的な遺伝子改変動物の開発にも取り組んでいます。これらを駆使することで、行動の多様性に関わる遺伝子を同定し、その機能を分子、細胞、更には神経レベルで明らかにすることを目指しています。
出版物
- Nakamura M, Nomoto K, Mogi K, Koide T, Kikusui T. Visual and olfactory signals of conspecifics induce emotional contagion in mice. Proc Biol Sci. 2024 Dec;291(2036):20241815.
- Niimura Y, Biswa BB, Kishida T, Toyoda A, Fujiwara K, Ito M, Touhara K, Inoue-Murayama M, Jenkins SH, Adenyo C, Kayang BB, Koide T. Synchronized Expansion and Contraction of Olfactory, Vomeronasal, and Taste Receptor Gene Families in Hystricomorph Rodents. Mol Biol Evol. 2024 Apr 2;41(4):msae071.
- Venkatachalam B, Biswa BB, Nagayama H, Koide T. Association of tameness and sociability but no sign of domestication syndrome in mice selectively bred for active tameness. Genes Brain Behav. 2024 Feb;23(1):e12887.
- Takanami K, Kuroiwa M, Ishikawa R, Imai Y, Oishi A, Hashino M, Shimoda Y,Sakamoto H, Koide T. Function of gastrin-releasing peptide receptors in ocular itch transmission in the mouse trigeminal sensory system. Front Mol Neurosci. 2023 Nov 30;16:1280024.
