生態遺伝学研究室
北野研究室
適応放散の遺伝機構
教員


Research Summary
どうやって新たな種が生まれるのか。生き物がどのようにして多様な環境に適応していくのか。生物多様性進化を巡るこれらの問いに対して、トゲウオやメダカを用いながら迫ります。表現型変化に関わる遺伝子は、実験モデル生物において多く同定されてきましたが、野外生物における種分化や適応進化の分子機構は多くが未解明です。また、原因対立遺伝子が野外集団内でどのように広まっていくのかについても多くが未解明です。これらを解明するために、フィールド調査から始まり、ゲノミックスや遺伝子工学、生態実験などを統合的に用います。
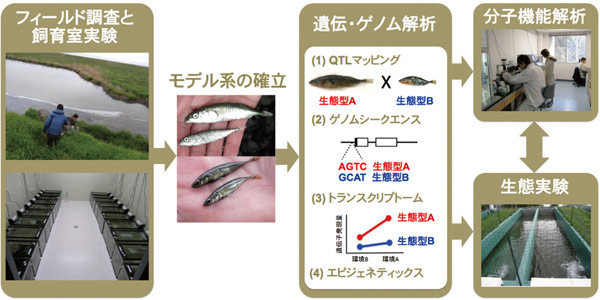
ついで、古典的な遺伝的手法や最新のゲノミックス技術などを援用し、遺伝基盤や候補遺伝子を解明していきます。また、遺伝子操作法による分子機能解析に加え、池やメソコスムを用いたミクロ生態系での生態実験を用いて分子から生態までをつないでいきます。
出版物
- Kitano J, Ansai S, Fujimoto S, Kakioka R, Sato M, Mandagi IF, Sumarto BKA,Yamahira K. A Cryptic Sex-Linked Locus Revealed by the Elimination of a Master Sex-Determining Locus in Medaka Fish. Am Nat. 2023 Aug;202(2):231-240.
- Yoshida K, Kitano J. Tempo and mode in karyotype evolution revealed by a probabilistic model incorporating both chromosome number and morphology. PLoS Genet. 2021 Apr 16;17(4):e1009502.
- Ansai S, Mochida K, Fujimoto S, Mokodongan DF, Sumarto BKA, Masengi KWA,Hadiaty RK, Nagano AJ, Toyoda A, Naruse K, Yamahira K, Kitano J. Genome editing reveals fitness effects of a gene for sexual dichromatism in Sulawesian fishes. Nat Commun. 2021 Mar 1;12(1):1350.
- Ravinet M, Kume M, Ishikawa A, Kitano J. Patterns of genomic divergence and introgression between Japanese stickleback species with overlapping breeding habitats. J Evol Biol. 2021 Jan;34(1):114-127.
