細胞建築研究室
木村研究室
細胞建築学:細胞の定量観察と理論モデルをとおして「建築家のいない建築」を理解する
教員


Research Summary
細胞は自然が造りあげたみごとな建築物です。細胞はその内部で、適切なサイズで形成された細胞内小器官が適材適所に配置していますが、この調和は中枢からの指令に基づくものではなく、多くの分子が自己組織的に作り上げています。細胞建築研究室では、「『建築家のいない建築』がどのように構築されているか?」をテーマに、細胞の顕微鏡観察と定量化、理論モデリングなどの手法を駆使し、細胞内の物質移動(オルガネラ配置や細胞質流動)やそのサイズ制御、隣接する細胞同士の協調による形づくりの研究をすすめています。
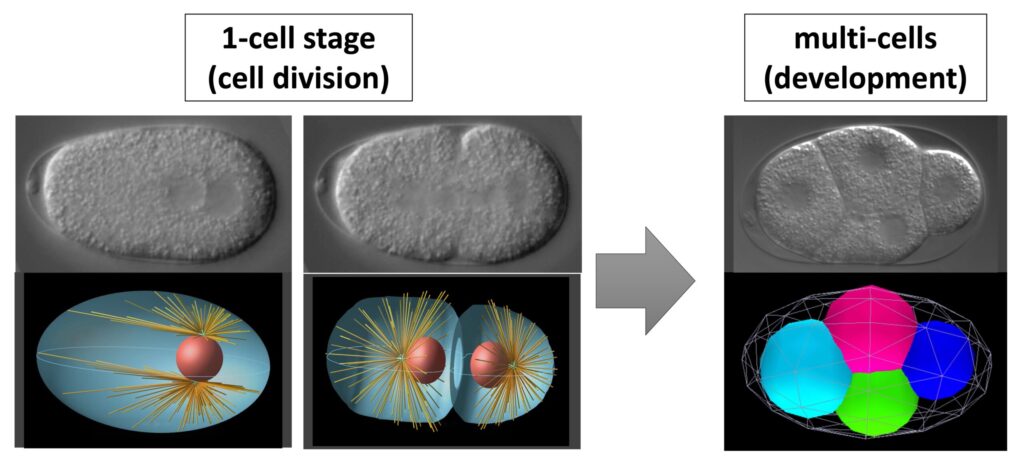
上段は実際の胚の顕微鏡像で、下段は定量的シミュレーション。
(右下の細胞配置の計算結果は慶應義塾大学・舟橋啓博士開発のソフトウェアを用いて表示したもの。)
出版物
- Ishikawa T, Torisawa T, Wada H, Kimura A. Swirling instability mediated by elastic and hydrodynamic couplings in cytoplasmic streaming. PRX Life. 2025.3, 013008.
- Goda M, Shribak M, Ikeda Z, Okada N, Tani T, Goshima G, Oldenbourg R, Kimura A. Live-cell imaging under centrifugation characterized the cellular force for nuclear centration in the Caenorhabditis elegans embryo. Proc Natl Acad Sci U S A. 2024 Oct 22;121(43):e2402759121.
- Kimura A. Quantitative Biology–A Practical Introduction. Springer 2022.
- 木村暁「細胞建築学入門」工学社2019.
