遺伝情報分析研究室
池尾研究室
ゲノム配列と遺伝子発現からみた分子進化学
教員

Research Summary
本研究室では、生物が新規の形質や特性を獲得するための分子基盤とその進化過程の解明を目指し、動物や菌類、細菌類を材料としてゲノム配列や遺伝子発現情報の比較解析を行っています。
特に
- 感覚器の進化に伴う遺伝子の分子進化解析
- メタゲノム解析を用いた微生物の多様性と環境ダイナミクスの解明
- ミトコンドリア及び核DNAに基づく分子系統解析
- データ解析による疾患原因遺伝子の探索と疾患モデルの構築
- 情報科学を用いた大規模データ解析システムの開発と知識発見
に力を注いで研究を行っております。
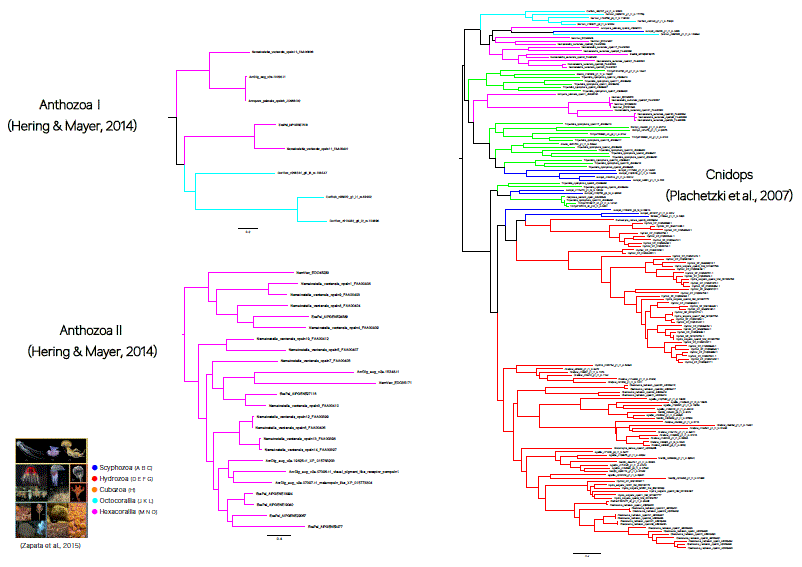
刺胞動物の持つオプシンは、大きく3つのグループに分かれ、且つそれぞれの系統内で独自に進
化してきたことが分かった。
出版物
- Ota N, Hirose H, Yamazaki Y, Kato H, Ikeo K, Sekiguchi J, Matsubara S, Kawakami H, Shiojiri N. Comparative study on a unique architecture of the brook lamprey liver and that of the hagfish and banded houndshark liver. Cell Tissue Res. 2024 Nov;398(2):93-110.
- Yoshida MA, Hirota K, Imoto J, Okuno M, Tanaka H, Kajitani R, Toyoda A, Itoh T, Ikeo K, Sasaki T, Setiamarga DHE. Gene Recruitments and Dismissals in the Argonaut Genome Provide Insights into Pelagic Lifestyle Adaptation and Shelllike Eggcase Reacquisition. Genome Biol Evol. 2022 Nov 4;14(11):evac140.
- Kinjo S, Kiyomoto M, Suzuki H, Yamamoto T, Ikeo K, Yaguchi S. TrBase: A genome and transcriptome database of Temnopleurus reevesii. Dev Growth Differ. 2022 May;64(4):210-218.
- Yuyama I, Higuchi T, Mezaki T, Tashiro H, Ikeo K. Metatranscriptomic Analysis of Corals Inoculated With Tolerant and Non-Tolerant Symbiont Exposed to High Temperature and Light Stress. Front Physiol. 2022 Apr 11;13:806171.
