Fluid-like behavior of chromatin in living human cell
Press release
Organization of fast and slow chromatin revealed by single-nucleosome dynamics
S. S. Ashwin, Tadasu Nozaki, Kazuhiro Maeshima, and Masaki Sasai
PNAS first published September 16, 2019 DOI:10.1073/pnas.1907342116
Press release (In Japanese only)
Understanding chromatin organization and dynamics is important, since they crucially affect DNA functions. In this study, we investigate chromatin dynamics by statistically analyzing single-nucleosome movement in living human cells. Bimodal nature of the mean square displacement distribution of nucleosomes allows for a natural categorization of the nucleosomes as fast and slow. Analyses of the nucleosome–nucleosome correlation functions within these categories along with the density of vibrational modes show that the nucleosomes form dynamically correlated fluid regions (i.e., dynamic domains of fast and slow nucleosomes). Perturbed nucleosome dynamics by global histone acetylation or cohesin inactivation indicate that nucleosome–nucleosome interactions along with tethering of chromatin chains organize nucleosomes into fast and slow dynamic domains. A simple polymer model is introduced, which shows the consistency of this dynamic domain picture. Statistical analyses of single-nucleosome movement provide rich information on how chromatin is dynamically organized in a fluid manner in living cells.
This research was supported by JST CREST(JPMJCR15G2), JSPS Kakenhi (JP19H05258, JP19H05273, JP19H01860, JP16H04746) and Takeda Science Foundation, RIKEN Pioneering Project、NIG Joint (2016-A2(6))
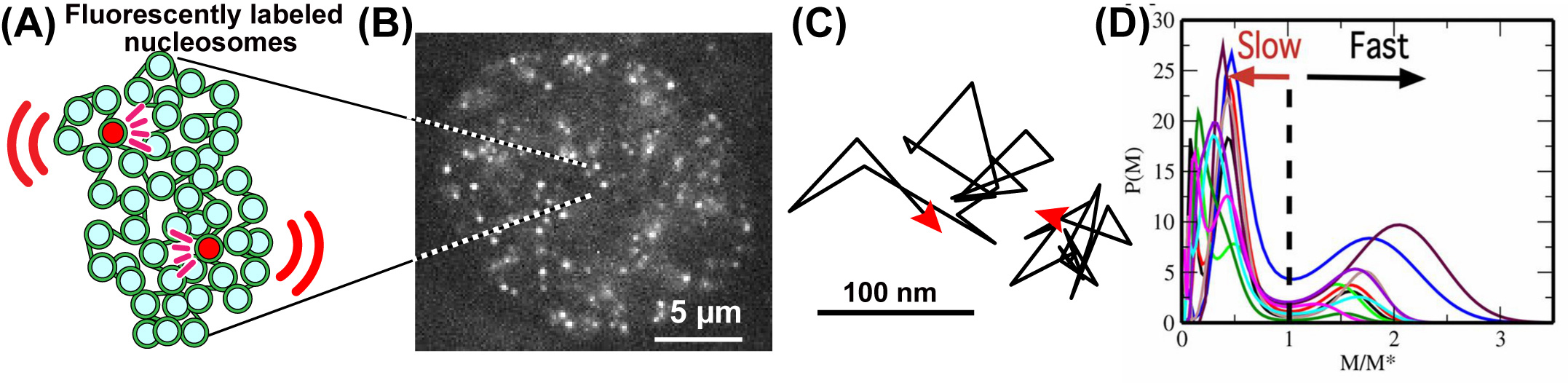
Fig: (A) A small fraction of nucleosomes, where DNA is wrapped around histone proteins, was fluorescently labeled (red). The labeled nucleosome movements can be tracked at super-resolution. (B) A single-nucleosome image of a living HeLa cell. (C) Representative two trajectories of the tracked single nucleosomes. (D) The distribution of MSD of single nucleosomes is plotted for 10-cell samples as functions of M/M*, where M* is M at the minimum between 2 peaks of the distribution.
Video: Raw video of single nucleosomes in the living HeLa cell. From Nozaki et al., (2017) Molecular Cell.















