A mechanism how intestine tumors affects liver functions
A novel zebrafish intestinal tumor model reveals a role for cyp7a1-dependent tumor-liver crosstalk in tumor’s adverse effects on host
Sora Enya, Koichi Kawakami, Yutaka Suzuki, Shinpei Kawaoka
Disease Models & Mechanisms (2018) DOI:10.1242/dmm.032383
The nature of host organs and genes that underlie tumor-induced physiological disruption on host has been poorly understood. Here, we establish a novel zebrafish intestinal tumor model, and find that hepatic cyp7a1, the rate-limiting factor for synthesizing bile acids (bile alcohol (BA) in zebrafish) is important for such a phenomenon. We created a transgenic zebrafish line, in which an oncogenic form of kras (krasG12D) was expressed in the posterior intestine by the Gal4-UAS method, and found that the intestinal tumor was formed. The intestinal tumor then caused detrimental effects on host, including liver inflammation, hepatomegaly, growth defects, and organismal death. Whole-organismal level gene expression analysis and metabolite measurements revealed that the intestinal tumor reduced total BA levels possibly via altered expression of hepatic cyp7a1. We demonstrated overexpression of cyp7a1 in the liver restored the BA synthesis and ameliorated tumor-induced liver inflammation. Thus, we discovered a previously unknown role of cyp7a1 as the host gene that links the intestinal tumor to the hepatic cholesterol-BA metabolism and liver inflammation. Our model provides an important basis to investigate host genes responsible for tumor-induced phenotypes and to uncover mechanisms underlying how tumors adversely affect host organisms.
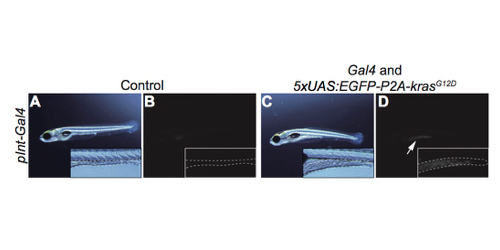
Figure: A transgenic zebrafish model bearing the intestine tumor created by the Gal4-UAS method. A, B; control. C, D: transgenic fish. B, D: fluorescent images. Dotted lines indicate the shape and size of the intestine. An arrow indicates EGFP expression.















