RNA sequences bound with a rice protein regulating meiosis transition
Experimental Farm • Nonomura Group
Rice MEL2, the RNA recognition motif (RRM) protein, binds in vitro to meiosis-expressed genes containing U-rich RNA consensus sequences in the 3′-UTR
Saori Miyazaki, Yutaka Sato, Tomoya Asano, Yoshiaki Nagamura, Ken-Ichi Nonomura
Plant Molecular Biology, Published online. 30 August, 2015. DOI:10.1007/s11103-015-0369-z
Developmental processes of plant germline are strictly regulated in angiosperms, to complete seed production during a short limited season. Synchronous male meiosis within a same anther, resulting in simultaneous production of pollen grains, is an example. Our group previously identified a rice protein, MEL2, regulating proper timing of meiosis transition. However, the mechanisms are largely ambiguous yet.
This paper identified RNA consensus sequences that MEL2 preferentially binds in vitro. Searched for the rice genome, the consensus-like sequences were frequently found in 3′-untranslated regions (3′-UTR) of 249 genes. Proteomic analyses revealed that several proteins that altered their amounts in mel2 mutant anthers were supposed to function in meiosis, and that the 3′-UTRs of their corresponding mRNAs contained MEL2-binding consensus-like sequences. These results suggest that MEL2 regulates meiosis transition via translational control of meiotic target mRNAs.
This is a collaborative work with National Institute of Agrobiological Sciences (NIAS) and Kanazawa University. This study was supported by JSPS KAKENHI and JSPS Bilateral Program.
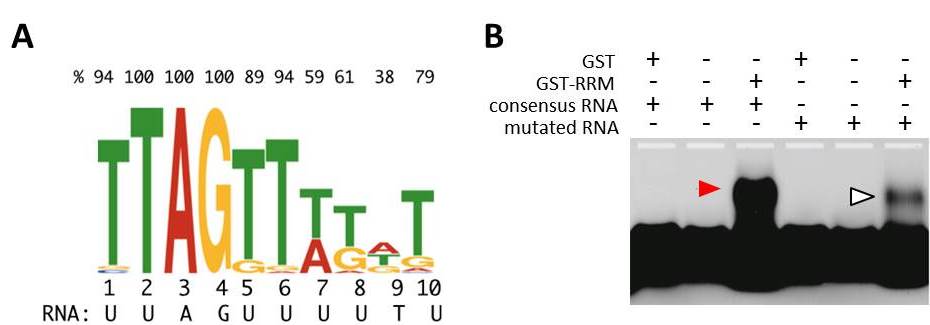
MEL2 protein binds uridine (U)-rich RNA sequences in vitro.
(A) 10-nucleotides RNA consensus sequence that RNA recognition motif (RRM) of MEL2 preferentially binds in vitro. Top values indicate the percentage of a prior residue (T, A, G or C) in each position.
(B) Electrophoresis mobility shift assay. The RNA-band position was shifted only when MEL2-RRM was mixed with consensus RNA sequence labeled with the fluorescense (red arrowhead). Mobility was significantly affected in the mutation-induced consensus (open arrowhead).















