物理細胞生物学研究室
島本研究室




生物物理の最先端技術による染色体動態制御機構の解明
教員

Research Summary
私達の体を構成する細胞の内部には核や紡錘体を初めとするミクロンサイズの構造体が多様に存在し、そのかたちや大きさをダイナミックに変化させながら染色体の動態を制御しています。当研究室では、生物物理に立脚した独自の顕微操作技術、一分子イメージング、試験管内再構成等の手法を駆使してこれらの構造体が示す時空間的変化を高解像で捉え、細胞分裂や胚発生の成功を支える分子機構を定量的に解明すべく研究を進めています。
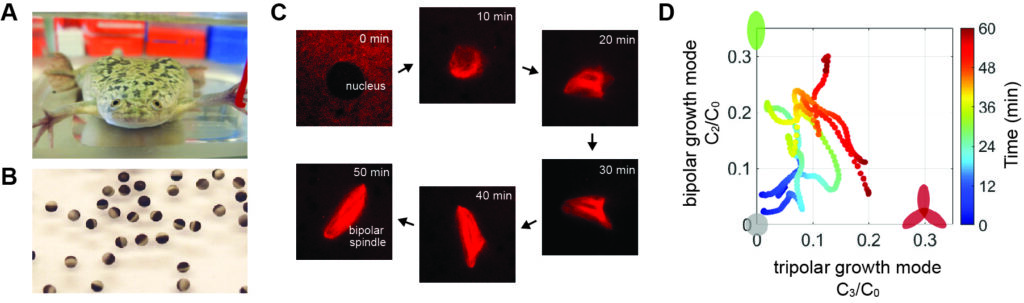
最近の研究では、ツメガエル卵(B)の細胞質抽出液を使った実験系と機械学習による画像解析手法を組み合わせ(C)、紡錘体と呼ばれる染色体分配装置が正しい二極性の構造へと自己組織形成されるしくみを明らかにした(D)。
出版物
- Sridhara A, Shimamoto Y. Microtubule choreography: spindle self-organization during cell division. Biophys Rev. 2024 Sep 30;16(5):613-624.
- Saju A, Chen PP, Weng TH, Tsai SY, Tanaka A, Tseng YT, Chang CC, Wang CH,Shimamoto Y, Hsia KC. HURP binding to the vinca domain of β-tubulin accounts for cancer drug resistance. Nat Commun. 2024 Oct 14;15(1):8844.
- Fukuyama T, Yan L, Tanaka M, Yamaoka M, Saito K, Ti SC, Liao CC, Hsia KC, Maeda YT, Shimamoto Y. Morphological growth dynamics, mechanical stability, and active microtubule mechanics underlying spindle self-organization. Proc Natl Acad Sci U S A. 2022 Nov;119(44):e2209053119.
- Mori M, Yao T, Mishina T, Endoh H, Tanaka M, Yonezawa N, Shimamoto Y, Yonemura S, Yamagata K, Kitajima TS, Ikawa M. RanGTP and the actin cytoskeleton keep paternal and maternal chromosomes apart during fertilization. J Cell Biol. 2021 Oct 4;220(10):e202012001.
