進化遺伝研究室
明石研究室
集団遺伝とゲノム進化
教員

Research Summary
我々の研究室はゲノム進化のメカニズム、特にゲノム中の大規模な適応進化と弱い自然選択の動態に焦点を当てて研究しています。機能ゲノミクスと進化理論に基づいて解析を計画し遂行することにより課題の解決に取り組んでいます。現在の研究テーマは以下の通りです。
- 同義変異にはたらく自然選択の表現型基盤と集団遺伝学
- アミノ酸組成・生合成がもたらすタンパク質進化への制約
- タンパク質の構造/機能と弱有害・適応進化の関係
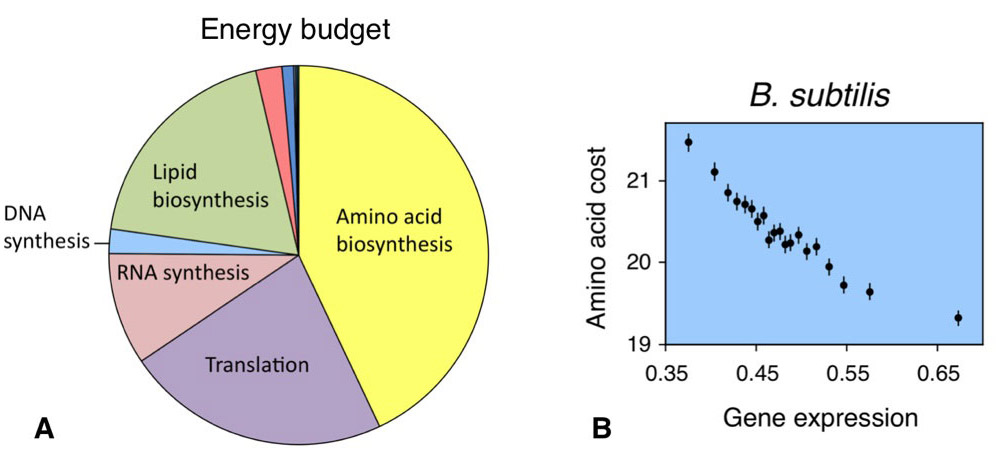
B)エネルギーコストに関わるタンパク質の進化。枯草菌のゲノムを見てみると、量の多いタンパク質はコストの安いアミノ酸を使って合成されていることが分かります。
出版物
- Matsumoto T, John A, Baeza-Centurion P, Li B, Akashi H. Codon Usage Selection Can Bias Estimation of the Fraction of Adaptive Amino Acid Fixations. Mol Biol Evol. 2016 Jun;33(6):1580-9.
- Matsumoto T, Akashi H, Yang Z. Evaluation of Ancestral Sequence Reconstruction Methods to Infer Nonstationary Patterns of Nucleotide Substitution. Genetics. 2015 Jul;200(3):873-90.
- Akashi H, Osada N, Ohta T. Weak selection and protein evolution. Genetics. 2012 Sep;192(1):15-31.
- Matsumoto T, Akashi H. Distinguishing Among Evolutionary Forces Acting on Genome-Wide Base Composition: Computer Simulation Analysis of Approximate Methods for Inferring Site Frequency Spectra of Derived Mutations. G3 (Bethesda). 2018 May 4;8(5):1755-1769.
