60-year scientific mystery of ’the DNA replication-timing program’ has finally been solved!
Replication timing maintains the global epigenetic state in human cells
Kyle N Klein, Peiyao A Zhao, Xiaowen Lyu, Takayo Sasaki, Daniel A Bartlett, Amar M Singh, Ipek Tasan, Meng Zhang, Lotte P Watts, Shin-Ichiro Hiraga, Toyoaki Natsume, Xuemeng Zhou, Timour Baslan, Danny Leung, Masato T Kanemaki, Anne D Donaldson, Huimin Zhao, Stephen Dalton, Victor G Corces, David M Gilbert
Science 372, 371-378 (2021) DOI:10.1126/science.aba5545
The temporal order of DNA replication is conserved from yeast to humans, but its biological significance remains unclear. We eliminated the protein RIF1, a master regulator of replication timing, in several human cell lines. RIF1 loss during the G1 phase of the cell cycle resulted in a heterogeneous, nearly random replication timing program from the first S phase. Altered replication timing was followed by replication-dependent redistribution of active and repressive histone modifications and alterations in genome architecture. These results support a model in which replication timing orchestrates the epigenetic state of newly replicated chromatin.
The Kanemaki laboratory contributed to the generation of a conditional RIF1 mutant using the auxin-inducible degron system.
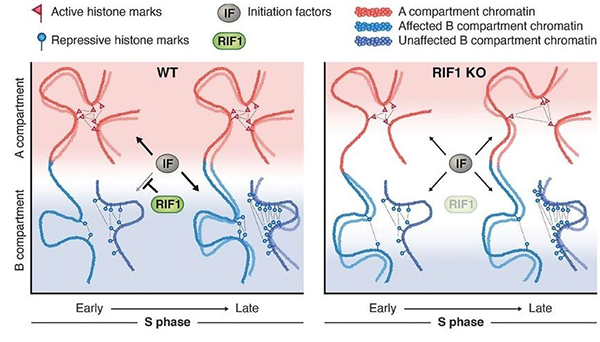
Figure: A model showing the role of the replication-timing program for the maintenance of the epigenome.
- You can read the “EurekAlert!” article here.















