A simple and efficient multiple sgRNA expression system in zebrafish
A tRNA-based multiplex sgRNA expression system in zebrafish and its application to generation of transgenic albino fish.
Tomoya Shiraki and Koichi Kawakami
Scientific Reports 8, Article number: 13366 (2018) DOI:10.1038/s41598-018-31476-5
The CRISPR/Cas9 system can be introduced into zebrafish fertilized eggs by microinjection and facilitates gene knock-out, genome editing, and gene knock-in studies in zebrafish. Further, expression of gRNA and controlled expression of Cas9 in transgenic fish have enabled the study of gene functions in specific cell types in zebrafish. The transgenic Cas9 approach would be more useful if multiple sgRNAs could be expressed simultaneously since we could knock-out a gene more efficiently or disrupt multiple genes simultaneously. However, such a system had not been developed in zebrafish.
In this study, we describe a novel system to express multiple sgRNAs efficiently in zebrafish, that relies on the endogenous tRNA processing machinery. In this study, we demonstrated that active sgRNAs can be produced from tRNA-fused precursor transcripts. To show a proof of principle, we constructed a transgenic fish line expressing a single transcript containing three distinct sgRNAs, that targeted the slc45a2 (albino) gene. We found that the transgenic-albino fish showed nearly complete albino phenotypes, which were transparent enough for imaging at the cellular resolution to be performed. Thus, the tRNA-based multiplex sgRNA expression system should facilitate genetic studies of gene functions in zebrafish.
This study was supported by JSPS KAKENHI Grant Numbers (JP15H02370, JP16K20983, JP16H01651, JP18H04988), and NBRP and NBRP/Genome Information upgrading program from Japan Agency for Medical Research and Development (AMED).
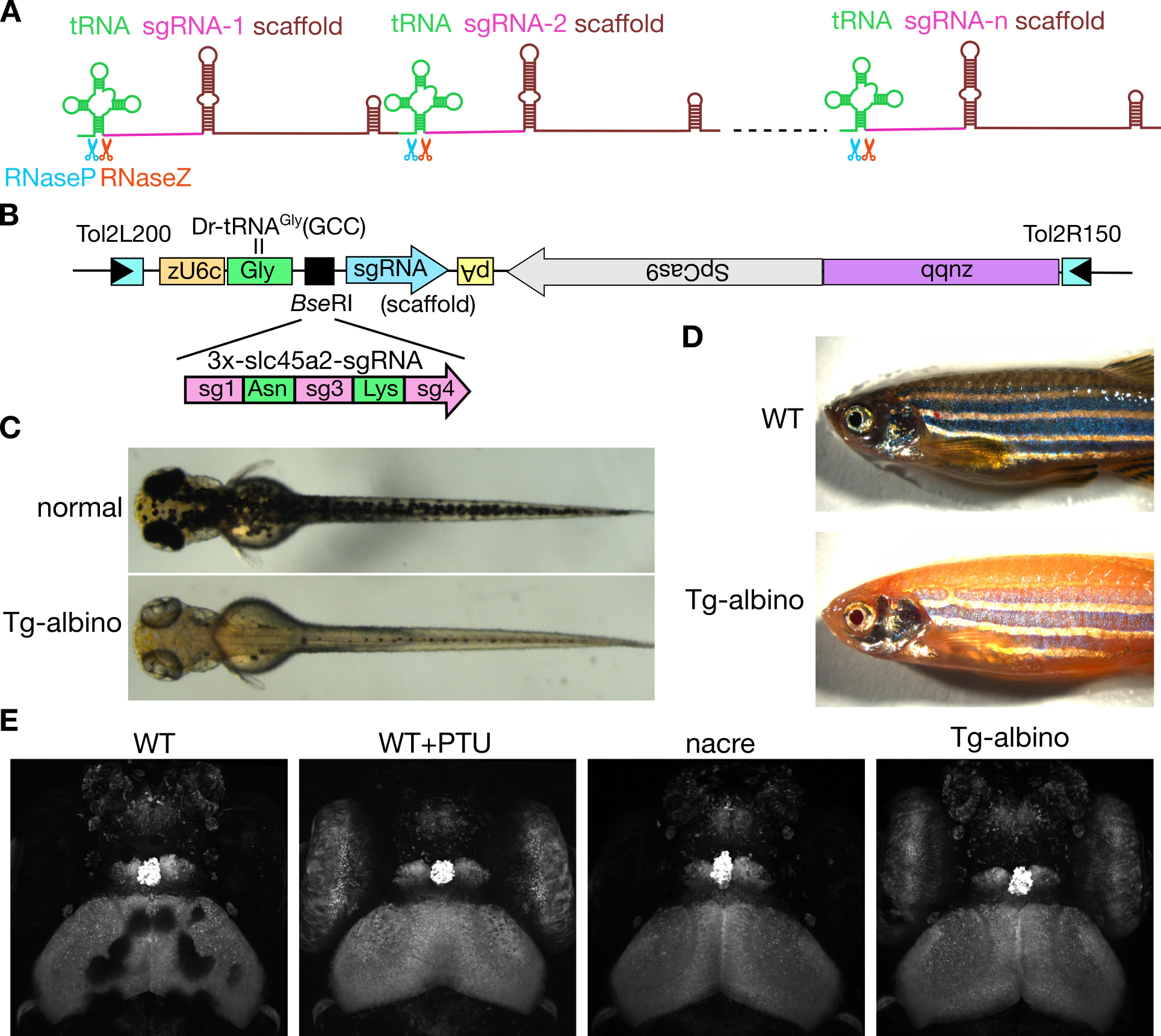
Figure: (A) Schematic diagram of the polycistronic tRNA-sgRNA system for multiplex genome editing. (B) DNA construct used for the generation of transgenic albino (Tg-albino) zebrafish. Three distinct sgRNAs targeting the albino (slc45a2) locus were expressed. (C-E) The Tg-albino fish showed nearly complete albino phenotypes, which were transparent enough for imaging at the cellular resolution. (C) Tg-albino at 3-days post-fertilization (dpf). (D)Adult Tg-albino fish. (E) Fluorescent images of a GFP-transgenic fish at 5 dpf in Tg-albino background.















