PZLAST-MAG: full length protein sequence similarity search server of large-scale MAG proteins
Koichi Higashi, Hitoshi Ishikawa, Ken Kurokawa, Hiroshi Mori
Bioinformatics Advances(2026)DOI:10.1093/bioadv/vbag129
A research group consisting of Associate Professor Hiroshi Mori and Professor Ken Kurokawa of the National Institute of Genetics, Research Organization of Information and Systems; Associate Professor Koichi Higashi of the AI-Empowered Life Science Initiative, Joint Support-Center for Data Science Research; and PEZY Computing K.K. has developed and freely released PZLAST-MAG, a web server that enables ultra-fast sequence similarity searches against approximately 400 million protein sequences (0.1 trillion amino acids) derived from over 210,000 metagenome-assembled genomes (MAGs) in the MAG database Microbiome Datahub. Implemented on PEZY-SC3 MIMD many-core processors, PZLAST-MAG delivers accuracy comparable to widely used tools such as DIAMOND and MMseqs2, while completing large-scale searches in just 5–15 minutes. Beyond standard tabular output, it offers interactive visualizations of phylogenetic distributions, environmental contexts based on the MEO ontology, and co-occurrence patterns of homologous proteins across MAGs (Figure 1). Use cases including CO₂-fixation marker enzymes and gut-associated plasmid proteins demonstrate its utility for homolog mining across diverse microbial lineages and ecosystems. PZLAST-MAG is freely available without registration at https://pzlast.nig.ac.jp/pzlast/mag.
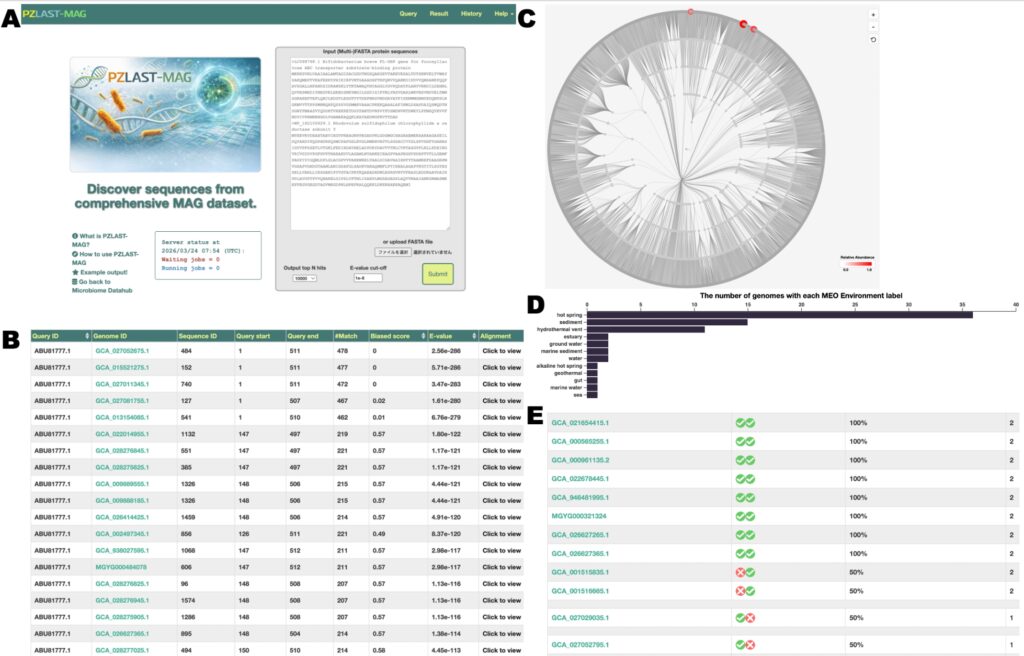
Overview of the PZLAST-MAG web server. (A) Search interface. (B) Tabular display of search results. (C) Display of results on a phylogenetic cladogram. (D) Environmental distribution of the hit MAGs. (E) Display of co-occurrence patterns across MAGs when multiple query sequences are submitted.
