Chromosome Biochemistry Laboratory
Murayama Group
Uncovering the molecular mechanisms underlying chromosome function through biochemical reconstitution.
Faculty


Research Summary
Controlling chromosome structure is essential not only for faithful chromosome segregation but also for gene transcription and DNA replication and repair. Ring-shaped SMC complexes (cohesin, condensin and SMC5/6) are central architects of the chromosome structure. These large complexes topologically entrap DNA strands to allow vital chromosomal functions to be carried out. We have successfully purified the SMC1/3 complex and reconstituted its functional DNA binding reaction. Our aim is to investigate the molecular mechanisms by which SMC complexes regulate the chromosome structure.
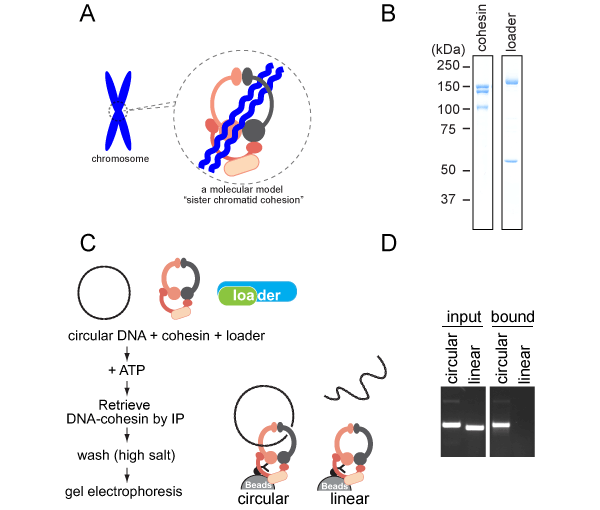
B. Purified cohesin proteins.
C, D. Biochemical reconstitution of topological DNA loading by the cohesin ring.
Selected Publications
- Murayama Y. Sister chromatid cohesion through the lens of biochemical experiments. Curr Opin Cell Biol. 2025 Jan 28;93:102464.
- Murayama Y, Endo S, Kurokawa Y, Kurita A, Iwasaki S, Araki H. Coordination of cohesin and DNA replication observed with purified proteins. Nature. 2024 Feb;626(7999):653-660.
- Kurokawa Y, Murayama Y. DNA Binding by the Mis4Scc2 Loader Promotes Topological DNA Entrapment by the Cohesin Ring. Cell Rep. 2020 Nov 10;33(6):108357.
- Murayama Y, Samora CP, Kurokawa Y, Iwasaki H, Uhlmann F. Establishment of DNA-DNA Interactions by the Cohesin Ring. Cell. 2018 Jan 25;172(3):465-477. e15.
