Genome Dynamics Laboratory
Maeshima Group
3D-organization and dynamics of human genome
Faculty

Research Summary
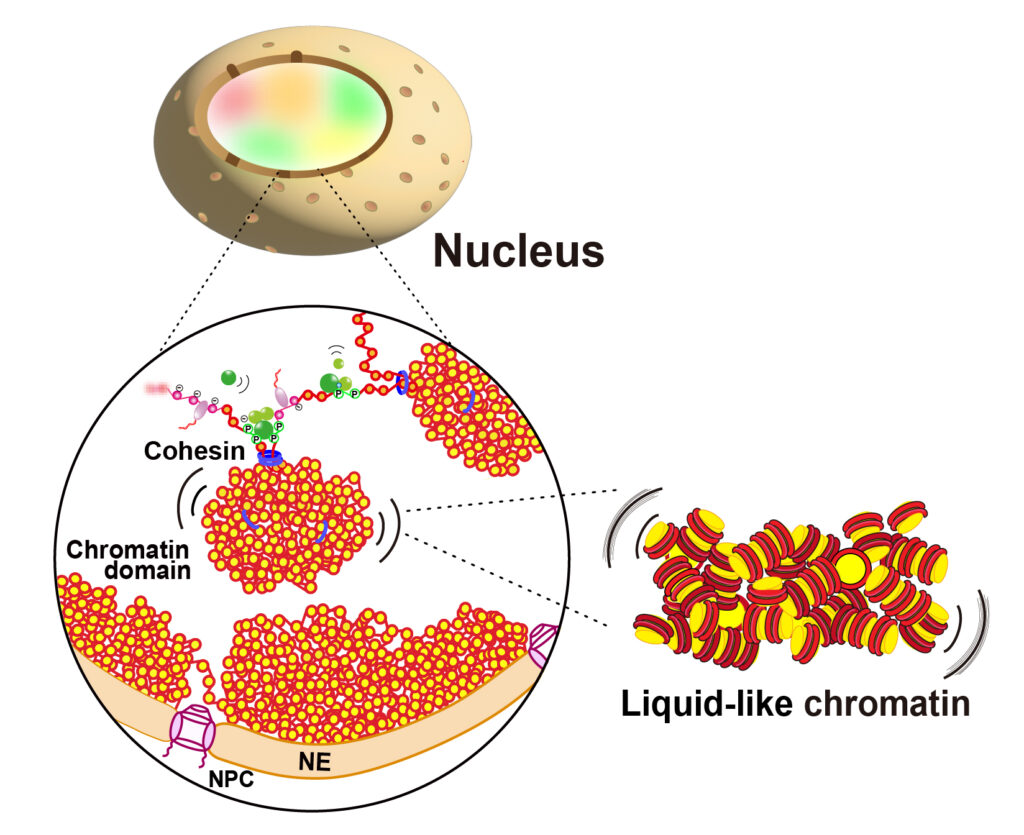
Our research interest is to know how a long string of human genome is three-dimensionally organized in the cell, and how the human genome is read out for cellular proliferation, differentiation and development. For this purpose, we are using a unique combination of molecular cell biology and biophysics, such as live-cell single molecule imaging, superresolution imaging, genomics and computational simulation.
Selected Publications
- Minami K, Nakazato K, Ide S, Kaizu K, Higashi K, Tamura S, Toyoda A, Takahashi K, Kurokawa K, Maeshima K. Sci Adv. 2025 Mar 28;11(13):eadu8400. doi: 10.1126/sciadv.adu8400.
- Iida S, Ide S, Tamura S, Sasai M, Tani T, Goto T, Shribak M, Maeshima K. Orientation-independent-DIC imaging reveals that a transient rise in depletion attraction contributes to mitotic chromosome condensation. Proc Natl Acad Sci U S A. 2024 Sep 3;121(36):e2403153121.
- Hibino K, Sakai Y, Tamura S, Takagi M, Minami K, Natsume T, Shimazoe MA,Kanemaki MT, Imamoto N, Maeshima K. Single-nucleosome imaging unveils that condensins and nucleosome-nucleosome interactions differentially constrain chromatin to organize mitotic chromosomes. Nat Commun. 2024 Aug 21;15(1):7152.
- Maeshima K, Iida S, Shimazoe MA, Tamura S, Ide S. Is euchromatin really open in the cell? Trends Cell Biol. 2024 Jan;34(1):7-17.
