Linker histone H1 functions as a liquid-like glue to organize chromatin in living human cells
Masa A. Shimazoe, Jan Huertas#, Charles Phillips#, Satoru Ide, Sachiko Tamura, Stephen Farr, S. S. Ashwin, Masaki Sasai, Rosana Collepardo-Guevara*, Kazuhiro Maeshima* #equal contribution; *corresponding authors
Science Advances(2026)DOI:10.1126/sciadv.aec9801
The paper was featured on the cover of the April 10 issue of Science Advances.
DNA inside the nucleus is not packed as a rigid regular fiber—linker histone H1 dynamically binds and loosely “glues” nucleosomes together, creating a dynamic, fluid organization that can still support essential genome functions.
The human genome has a remarkable capacity for compaction. If all 46 human chromosomes were stretched end to end, they would collectively reach about two meters in length, yet they are somehow arranged within a nucleus only ~10 micrometers in diameter. To fit inside the nucleus, DNA is wrapped around histone proteins, like thread around a spool, to form nucleosomes. Nucleosomes and associated proteins then fold into chromatin, enabling the genome to be packaged while remaining functional.
For decades, molecular biology textbooks described linker histone H1 as a key factor that organizes nucleosomes into regular, rigid higher-order structures (often depicted as a 30-nm chromatin fiber). However, accumulating evidence suggests that such highly ordered fibers are very rare in living cells. This raises a fundamental question: how does H1 actually compact DNA and organize chromatin in vivo?
A team led by Professor Kazuhiro Maeshima at SOKENDAI and the National Institute of Genetics (Research Organization of Information and Systems, ROIS) in Japan and Professor Rosana Collepardo-Guevara at the University of Cambridge in U.K. investigated how H1 binds to nucleosomes and shapes chromatin inside living human cells. By combining single-nucleosome imaging with super-resolution fluorescence microscopy and computational modeling, the researchers reveal a new physical picture of H1 function: rather than working as a static clamp (see Figure, left), H1 behaves like a dynamic, liquid-like “glue” that gently folds chromatin into a fluid organization that remains accessible to other molecules (Figure, right).
This research was supported by Japan Society for the Promotion of Science (JSPS) and MEXT KAKENHI grants JP23K17398, JP24H00061, JP22H00406, JP21H02535, and JP22H05606; the Takeda Science Foundation; the UK Government’s Guarantee scheme (EP/Z002028/1) following funding from the European Research Council (Consolidator Grant); the UK High-End Computing Consortium for Biomolecular Simulation (EP/R029407/1); the Herchel Smith Postdoctoral Fellowship; UK Research and Innovation (UKRI) Postdoctoral Fellowships Guarantee scheme (EP/X02332X/1); JSPS Research Fellowship JP24KJ1161; and the National Institute of Genetics 2023 NIG-JOINT (4I2023).
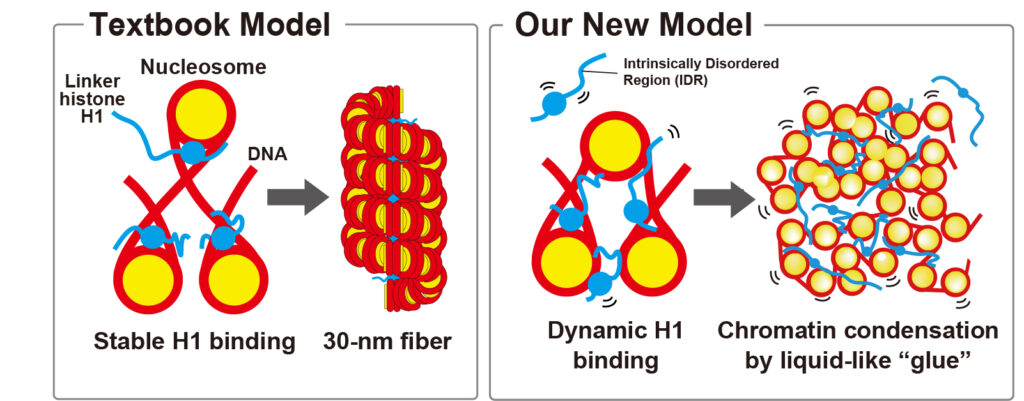
Previously, linker histone H1 was thought to bind stably to nucleosomes and compact chromatin by forming an ordered, “rigid” structure (left). In this study, we show that H1 binds to nucleosomes dynamically and acts like a liquid-like “glue,” compacting chromatin while creating a dynamic, fluid-like organization (right).
(Left) Video of linker histone H1 in the nuclei of human living cells observed by super-resolution microscopy. (Right) Molecular dynamics simulation reproducing interactions between nucleosomes and linker histone H1. Red, DNA; yellow, histones; blue, linker histone H1. H1 moves dynamically while bridging multiple nucleosomes.
