Cutting-edge Research : A Core Institute for Life Sciences
Life is a complex system generated by the mutual interaction between genetic information engraved in the genome and the internal and external environments. At the National Institute of Genetics (NIG), cutting-edge re-search is conducted in areas such as cell function, development and differ-entiation, evolution and diversity, and genome information, aiming to clarify the system of life.
Research Highlights
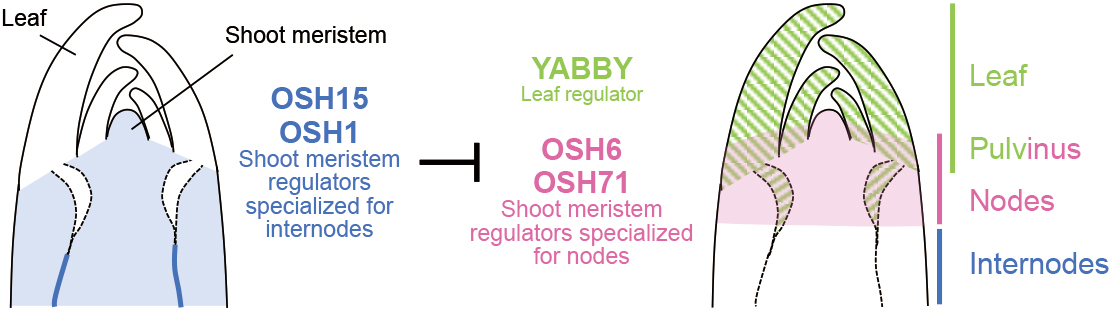
Development and evolution of plant stem nodes and internodes
The stem is the plant axis important in crop breeding to prevent lodging. Tsuda group showed that nodes and internodes, building units of the stem, are specified by regulators for shoot meristems and leaves, offering future avenues to manipulate crop height.
- Tsuda K, Maeno A, Otake A, Kato K, Tanaka W, Hibara KI, Nonomura KI. YABBY and diverged KNOX1 genes shape nodes and internodes in the stem. Science. 2024 Jun 14;384(6701):1241-1247.
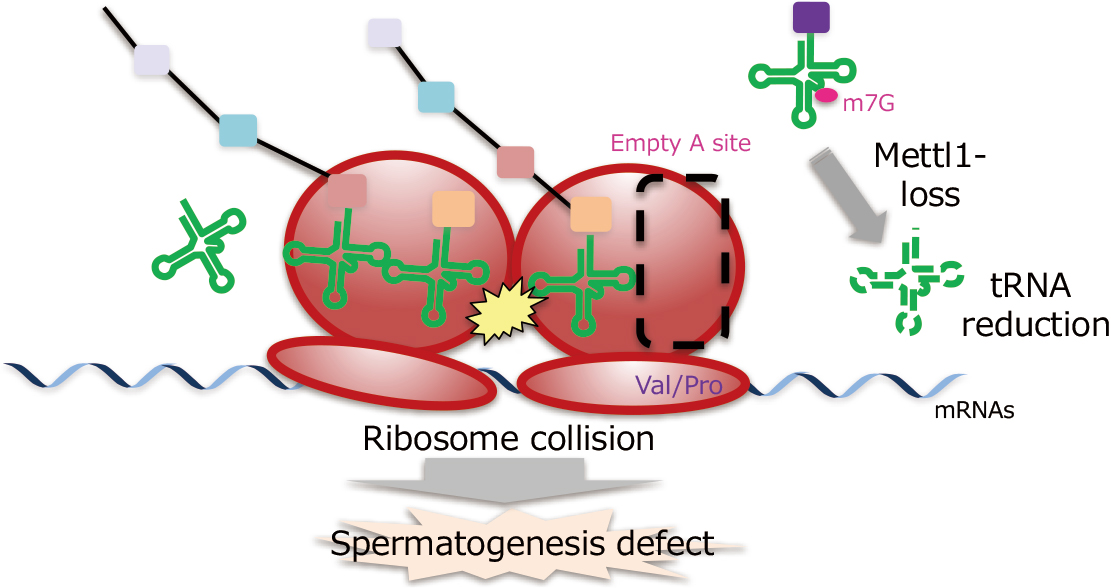
Physiological functions of tRNA m7G modification
Saito group discovered that Drosophila Mettl1 ensures the amount of tRNA and pro-
tein translation necessary for spermatogenesis by introducing a chemical modification
called m7G (N7-methylguanosine) into a subset of tRNAs. This research will contribute to
the understanding of cancer and neurological diseases that are reported to be caused
by abnormalities in RNA modification.
- Kaneko S, Miyoshi K, Tomuro K, Terauchi M, Tanaka R, Kondo S, Tani N, Ishiguro KI, Toyoda A, Kamikouchi A, Noguchi H, Iwasaki S, Saito K. Mettl1-dependent m7G tRNA modification is essential for maintaining spermatogenesis and fertility in Drosophila melanogaster. Nat Commun. 2024 Sep 24;15(1):8147.
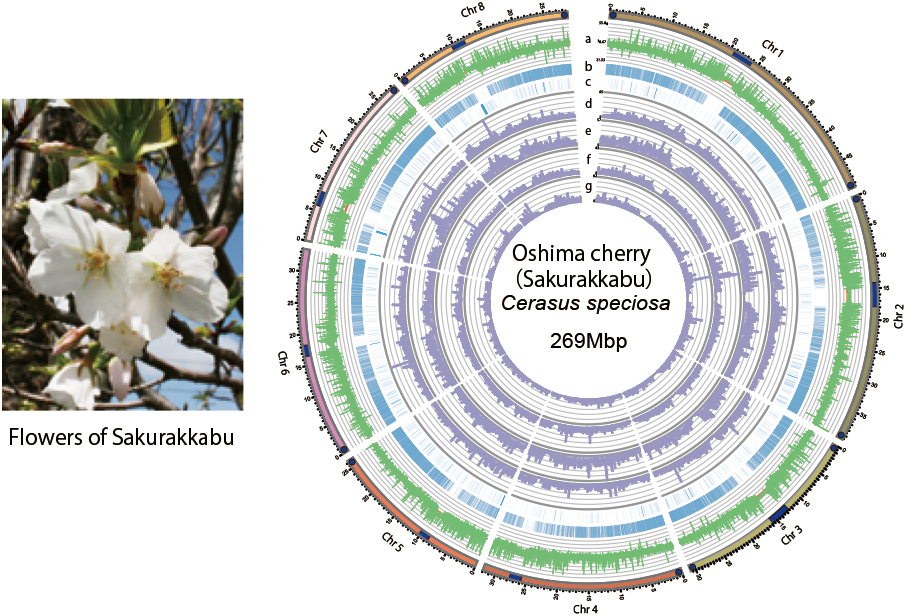
Complete genome sequence of Oshima cherry Cerasus speciosa released
The complete genome sequence of the Oshima cherry, an endemic species in Ja-pan, has been successfully decoded by Dr. Koide and a research team ‘Sakura 100 Ge- nome Consortium’. As a sample, leaves obtained from a Natural Monument of Japan,over 800 years old, located on Izu Oshima Island were used.
- Fujiwara K, Toyoda A, Biswa BB, Kishida T, Tsuruta M, Nakamura Y, Kimura N, Kawamoto S, Sato Y, Katsuki T; Sakura 100 Genome Consortium; Koide T. A Near Complete Genome Assembly of the Oshima Cherry Cerasus speciosa. Sci Data. 2025 Feb 4;12(1):162.