


Molecular evolution is a branch of biology that studies how organisms evolve based on molecular information including DNA and protein sequences. Professor Shigehiro Kuraku has been investigating the origins of biological phenomena, mainly in developmental biology, using DNA sequence analysis of diverse vertebrates. He states that “Molecular evolution is an essential tool for all fields of life science.”
Profile
Professor Kuraku graduated from the Faculty of Science at Kyoto University in 1999. After receiving his PhD from the Graduate School of Kyoto University in 2005, he served as a research scientist at RIKEN. He obtained a junior faculty position at the University of Konstanz (Germany) in 2007. He became the Unit Leader at RIKEN CDB in 2012 and became Team Leader, also at RIKEN BDR in 2019. He was appointed Professor at the National Institute of Genetics (NIG) in April 2021.
Shark research is critical to overview vertebrates
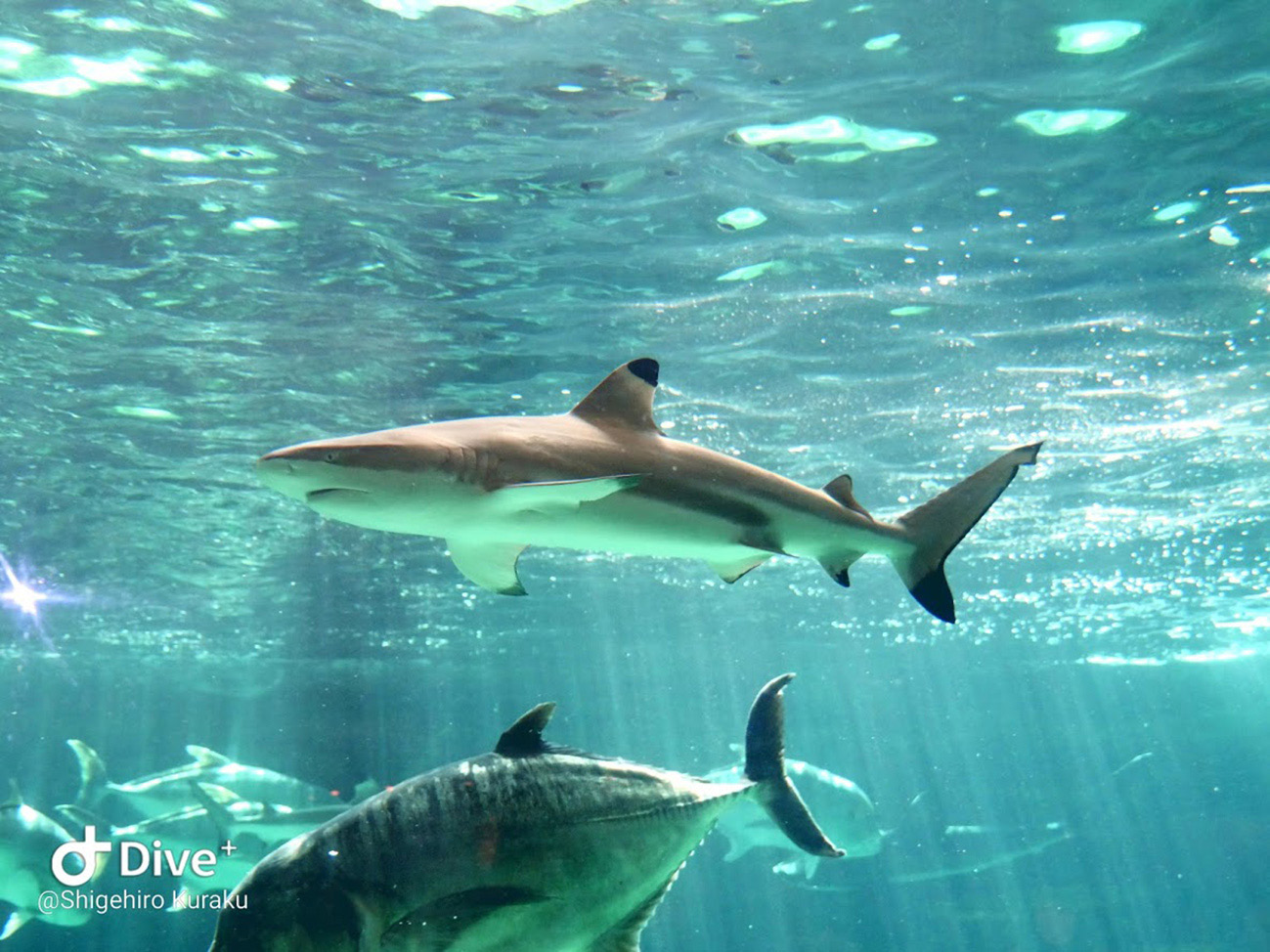


I am planning two directions for my research at NIG. First, I would like to analyze groups of animals for which there is not a lot of genome sequence information, in order to make them equally comparable with other groups. Second, after comparing genomes, I plan on finding any distinct differences and clarifying how these differences contribute to the characteristics that make up the identity of an organism. I would like to explain this by introducing some of my research. This includes research I carried out on sharks, which I published in 2018.
There were two groups of vertebrates that were severely lacking in detailed genome information: cartilaginous fish and reptiles. Cartilaginous fish mainly comprise sharks and rays. We analyzed the genomes of three species: the bamboo shark, the catshark, and the whale shark. Genome analysis of shark species was lagging behind partly because their genome is large and difficult to analyze. However, we were able to achieve a highly comprehensive analysis that no one else could match at the time. I think the work we carried out was a rarity in that it was completed in one laboratory from start to finish.
rarity in that it was completed in one laboratory from start to finish sharks. We also looked at organisms that have been well studied, such as humans, and clarified how their hormone repertoires are established.
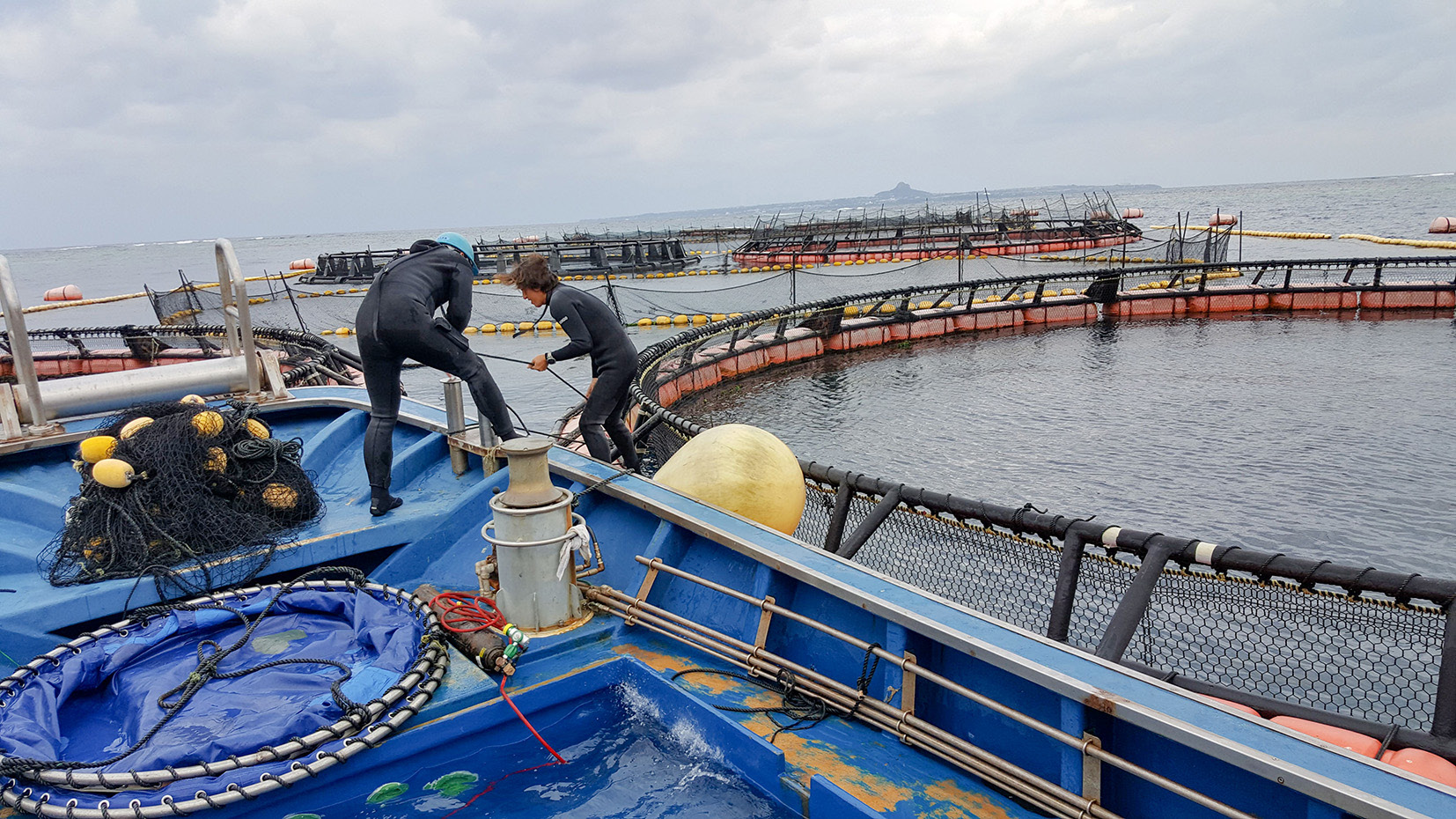


We also investigated opsin gene repertoires that control vision and found that the catshark may not see in colour. Nowadays, catsharks can reside in shallow water. However, our findings suggested that their ancestors were likely to have lived deep in the sea. We also revealed that a whale shark opsin gene can absorb light of wavelengths that can penetrate the deep sea. This knowledge regarding their deep-sea habitat was obtained through experiments where proteins were synthesized based on the genome sequences. This was all accomplished without the need to sacrifice living organisms.
The National Institute of Genetics is located close to Suruga Bay—a well-known deep-sea area. Here, we will continue to push forward with our research into sharks. Cartilaginous fish have a unique phylogenetic position among vertebrates, and yet little molecular research has been done on them. By analyzing these species, we will not only elucidate the evolutionary process of early vertebrates that existed on Earth 500 million years ago but also explore the conditions of ecological systems in the sea that need to be protected.
The research presented here includes the two directions I previously mentioned: organizing information that is the basis for comparison at a molecular level, and shedding light on biological phenomena. There is also another common characteristic behind my research methods that I would like to emphasize—a means to connect with different scientific disciplines through research on molecular evolution. I would like to explain this while introducing my career to you.
First encounter with molecular evolution
When I first learned about molecular evolution, I was still a high school student preparing for my university entrance exams. It was just one week before the National Center Test for University Admissions. By chance, I was drawn to a book in a bookstore authored by Professor Takashi Miyata of Kyoto University (Now professor emeritus of Kyoto University) called “Invitation to the Study of Molecular Evolution,” and ended up reading it from start to finish in one sitting. The following is a quotation from the book: “Conventional research on evolution was limited to comparing superficial phenomena, such as morphology, and tended to be subjective. However, the study of molecular evolution will make the research quantitative and objective for the first time.”
I entered Kyoto University where Professor Miyata conducted his research and attended his lectures at every opportunity. I had no reservations about joining Professor Miyata’s laboratory for my undergraduate research, and there I learned how to conduct research on molecular evolution.
In the study of molecular evolution, an evolutionary pathway is deduced after the inference of a phylogenetic tree based on DNA and protein sequences of biological species. To be specific, it is necessary to build a phylogenetic tree by comparing the sequences of a broad range of species. Since a vast number of calculations are required, you sometimes have to write your own programs and carry out calculations using a computer specialized for large-scale analysis. At the Miyata Laboratory, I was able to learn about molecular evolutionary analysis starting from the basics. This included using a computer to compare sequences, calculating the divergence time (age) of particular species, and building phylogenetic trees.
Taking the plunge into the world of developmental biology


When I was a graduate student, the availability of animal genome sequences was very limited. I sometimes performed experiments to clone gene fragments of the organisms I was interested in. However, most of my time was spent sitting in front of a computer analyzing the nucleotide sequences of the DNA, consisting of A, T, G, and C. Before long, I thought I should also train myself with hypothesis-based experiments using cells and tissues. That is what most other researchers were working on.
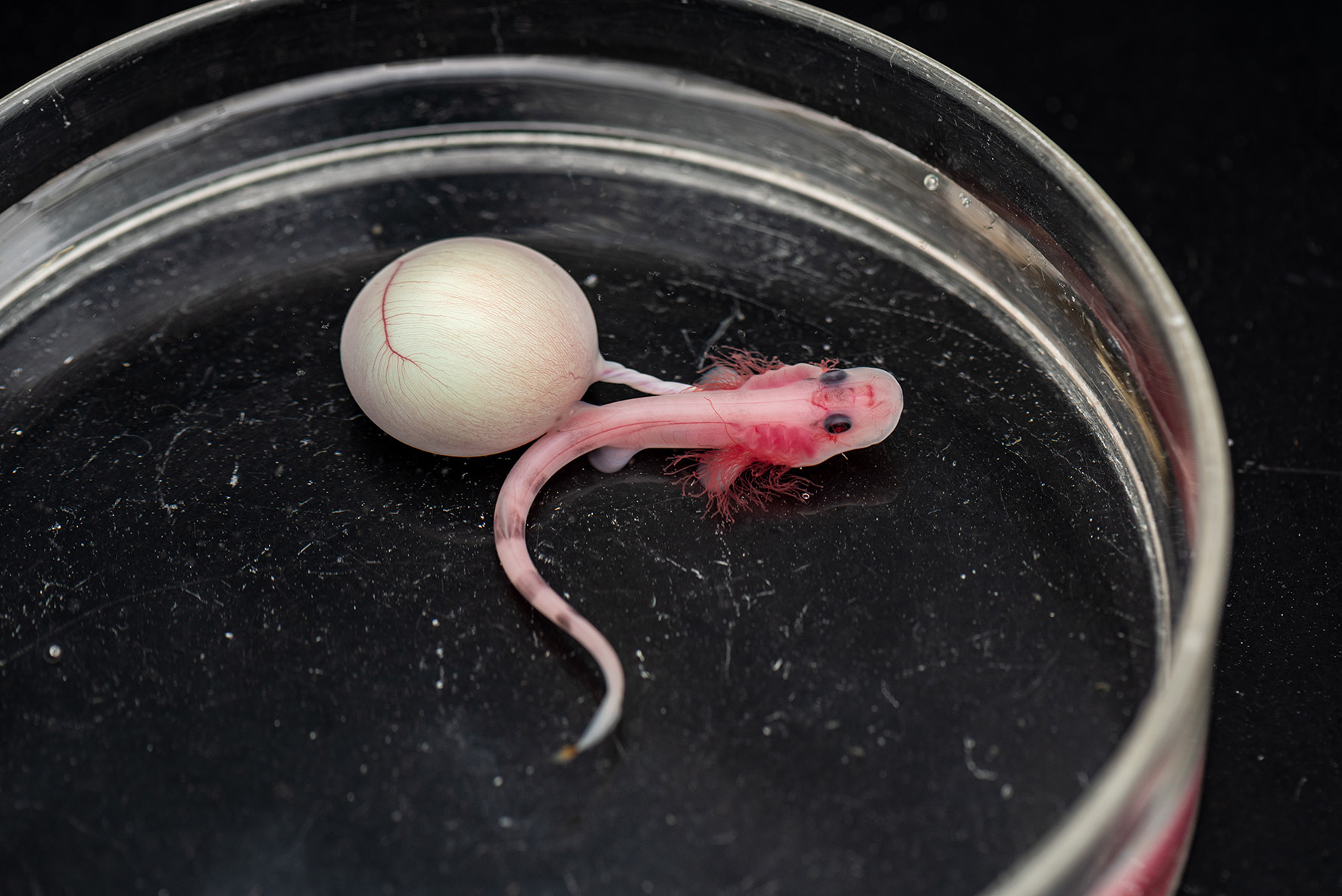
In my doctoral program, I decided to join a laboratory that was conducting research into developmental biology. From there, I commenced my own research in the laboratory of Dr. Shigeru Kuratani of the RIKEN CDB (RIKEN Center for Developmental Biology). The Kuratani laboratory specializes in comparative developmental biology studies, in particular, EvoDevo (which means “evolutionary developmental biology”). My research was focused on the evolution of the shells of soft-shelled turtles, and I obtained my PhD based on this research theme.
At that time, methods exploring functional changes of developmental regulatory genes while keeping in mind the evolution of those genes were not established. I was already familiar with molecular evolution and felt as though a fresh perspective had opened up in front of me. I realized that if I were to link developmental biology together with molecular evolution, I could play a role of my own in the field.
At the Kuratani Lab, I had many opportunities to get involved in other members’ research. I started considering how I could solve problems by applying my skills, and what was required to move the whole project forward. This experience trained me well and thereafter it became a fundamental part of my research style.
At the Kuratani Lab, I had many opportunities to get involved in other members’ research. I started considering how I could solve problems by applying my skills, and what was required to move the whole project forward. This experience trained me well and thereafter it became a fundamental part of my research style.
Conducting research based on my own plan
After I received my Ph.D., I still had some research to do, so I stayed at the Kuratani Lab for two more years as a postdoctoral researcher. I had already been looking for a postdoctoral position before I got my degree. My primary goal was to study abroad, but I guess I was lacking something since I missed out on all the opportunities I applied for. At this stage, I was aiming to get a fellowship and carry out research on my initiative, rather than getting paid to do lab work. I think this idea narrowed my opportunities to get accepted.
In my second year, I changed my mind and decided to look for PI (Principal Investigator) positions. I figured this was best if wanted to try working with my ideas. There was a public offering for a junior leader position at University of Konstanz in Germany which I thought was a perfect fit. It was posted on a job site run by Nature, and I felt it was an opportunity I couldn’t pass up.
Dr. Axel Meyer, a professor at the university who posted the position, specializes in evolutionary biology. By chance, I had met him at a mixer at an international conference just three months previously, where we sat down and had a conversation. I applied for the job immediately, and fortunately, I was hired.
I worked as a junior leader for five years at University of Konstanz, located near the Swiss border, starting in 2007. The University was selected as the target for a national excellence initiative project that year. As a junior leader, I was treated as a member of a large research group led by the professor and was responsible for undergraduate education including practical training courses in addition to general lectures. I was able to freely pursue research themes based on my own ideas, while communicating with the professor.
From the second year of my appointment, I applied for the German Research Foundation (DFG) grant, which is similar to the Japanese Grant-in-Aid for Scientific Research. As a result, I was successful in securing research funding. There are no categories based on age or experience, and applications are accepted year-round. I felt that this is very different from the Japanese system.
My time in Germany: preparing for lectures


This is how I was able to secure a position abroad where I could conduct research based on my own ideas. Initially, I spent all of my time preparing for teaching. Fortunately, I was in charge of a class where I could communicate in English, and not in German. Despite having an interest in language and being proficient in English, it was not easy to teach. First and foremost, I had no prior teaching experience even in Japanese, and it took more than two years before I was able to adjust and think through things calmly.
t was only after I had a few PhD students under my wing and could hire part-time students that I was able to focus on my research.
At that time, genome analysis of species other than conventional experimental animals was gaining traction. Taking advantage of this trend, I joined the International Genome Consortium of Sea Lamprey. I also got the opportunity to have a say in the overall direction of projects by producing most of the figures for the published paper. Through this experience, I was able to get a feel for the direction that genome analysis was going in around the world and could use this knowledge to my advantage in my own research. Using the sequence information developed by the Consortium, my research was mainly focused on the evolution of the gene repertoire that controls the development of vertebrates. This covered many vertebrates, including lampreys.
At that time, genome analysis of species other than conventional experimental animals was gaining traction. Taking advantage of this trend, I joined the International Genome Consortium of Sea Lamprey. I also got the opportunity to have a say in the overall direction of projects by producing most of the figures for the published paper. Through this experience, I was able to get a feel for the direction that genome analysis was going in around the world and could use this knowledge to my advantage in my own research. Using the sequence information developed by the Consortium, my research was mainly focused on the evolution of the gene repertoire that controls the development of vertebrates. This covered many vertebrates, including lampreys.
Having a job in data production in Japan
Around my fourth year at University of Konstanz, I started looking for my next position. I was particularly conscious about current trends, and up until that point I was using genome information prepared by others in my research. The continuity of genomic DNA sequences was often lacking, and in order to make my analyses more accurate, I figured it was necessary to become familiar with DNA sequencing technology—something that had already achieved huge progress at the time. As a result, I was hired for a leadership role by RIKEN CDB that was setting up a new division using this cutting-edge technology. So, after five years of life in Germany, I decided to move back to Japan.
The main role of the position at RIKEN was to support NGS (next-generation sequencer) analysis at the request of the researchers in the institute. I was a former employee of RIKEN, but this time around it was an entirely different position. I started my life thereby learning about new advanced technologies, putting off my research. In my nine years of service there, I gained extensive experience in analyzing different information, from sequencing genomes to profiling gene expression. Researchers from various fields of developmental biology were working around me, and I tried my best to fulfil the role of supporting DNA information analysis through a number of collaborative research projects. Using the experience I gained in my doctoral course, where I worked with the approach of incorporating different fields of study into developmental biology, I felt I was applying the same approach on a broader scale in my work.
The main role of the position at RIKEN was to support NGS (next-generation sequencer) analysis at the request of the researchers in the institute. I was a former employee of RIKEN, but this time around it was an entirely different position. I started my life thereby learning about new advanced technologies, putting off my research. In my nine years of service there, I gained extensive experience in analyzing different information, from sequencing genomes to profiling gene expression. Researchers from various fields of developmental biology were working around me, and I tried my best to fulfil the role of supporting DNA information analysis through a number of collaborative research projects. Using the experience I gained in my doctoral course, where I worked with the approach of incorporating different fields of study into developmental biology, I felt I was applying the same approach on a broader scale in my work.
Planning a new type of research on molecular evolution at NIG

I began to think that setting up laboratories focused on molecular evolution and providing guidance to young people was the responsibility of those who specialize in the field. I came to NIG with that idea. Today, I feel that molecular evolution is not only a single academic field, but also a valuable tool for all life science researchers. In the shark research I explained at the beginning of this interview, molecular evolutionary analysis techniques were used in discoveries related to developmental biology, comparative endocrinology, and ecology.
Many interesting animals have undergone striking phenotypic evolution. For example, whales live in the sea even though they are mammals, and turtles have evolved to have shells. In this regard, I would like to find out how the genome determines the characteristics of each organism by looking at a variety of organisms and having an overview of the entire genome. People may think that to study evolution is to study the past. However, to study evolution is to investigate how a living organism has evolved into what it is today and how it differs from other organisms that make up the ecosystem. In other words, it is also a study that teaches the “now” of biodiversity.
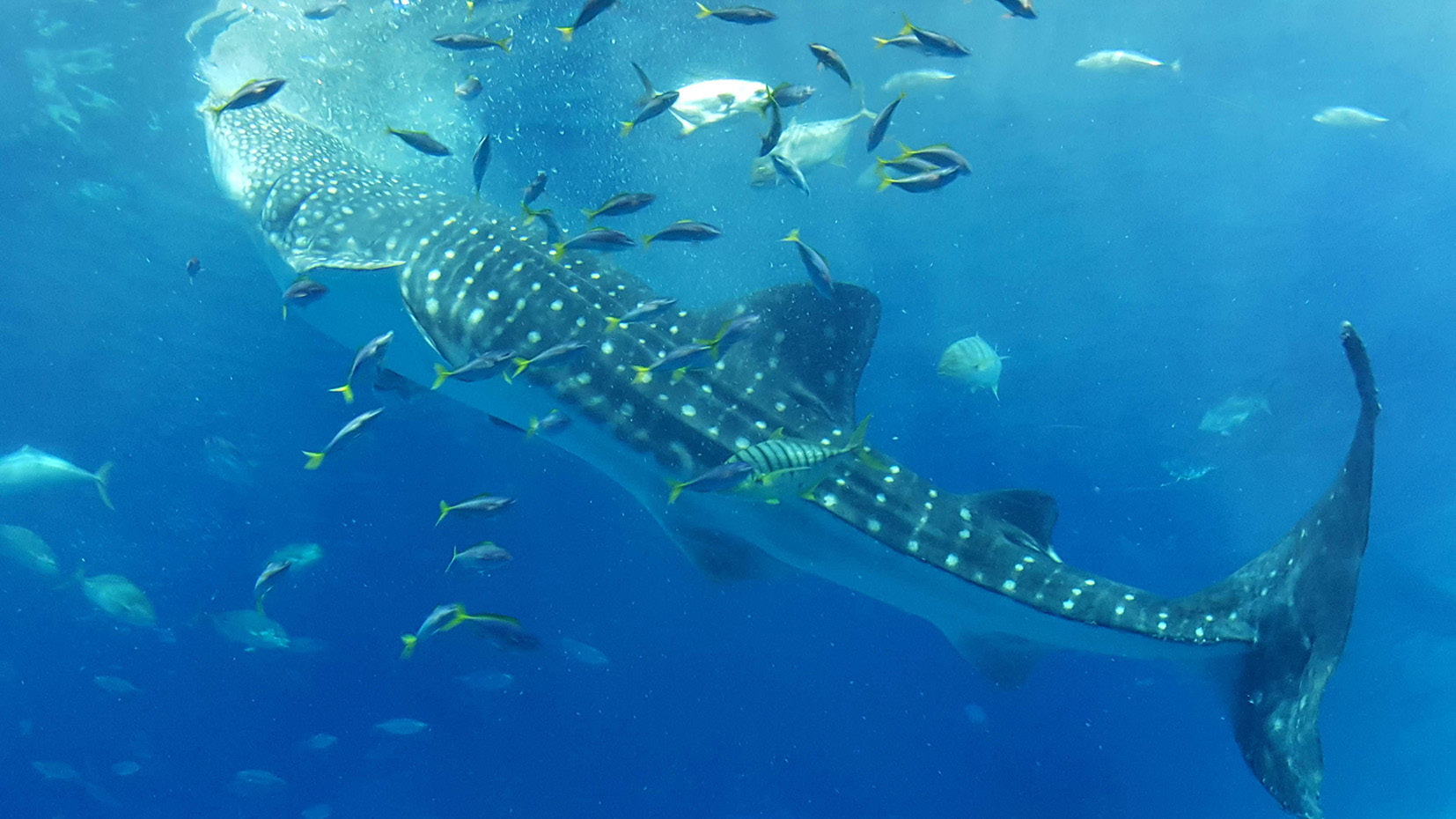


Another method of research I am focusing on is analysis at the cellular level. As technology advances, acquiring omics information at the single-cell level has become an increasingly active field around the world. These techniques determine how genome information is being expressed. Much as the key concept behind the Neutral Theory of Molecular Evolution is that there are functional constraints on gene functions, functional constraints to achieving cellular activity should also exist. However, our knowledge is limited to organisms that have short lives and small genomes. If we apply a molecular evolutionary approach to omics information, I think we will be able to come close to answering various questions, such as what are the factors that determine the size of the genome? —something we don’t yet have an answer for. It is important to compare and explore not only model organisms but also various organisms in nature, using a molecular evolutionary approach.
Message to young people


It is my motto not to narrow down my research subjects. Many researchers narrow down their themes and concentrate on a focused topic. This creates a gap between one research field and another. I always want to keep my eyes open and connect these gaps using an approach I’m good at. I think connecting fields and elements together is a process that helps push science forward, and it is something I am particularly enthusiastic about.
I think molecular evolution plays a role in making these connections. With this in mind, I believe it is critical to have a broad perspective. I named my laboratory the “Molecular Life History” Lab with the idea that we would not limit ourselves to a particular field or, conversely, exclude any other fields. When we try to elucidate the origins of life at a molecular level, which is very complex, we must face the problem head-on by incorporating techniques and knowledge from all research fields. Those interested in joining my laboratory are expected to learn how to develop a perspective that goes beyond a single field of study and to expand their networks for this purpose.
If you are going to be a researcher, my wish is that you develop your communication skills to engage in creative activities. It is important to have the ability to deepen your research through communication with others, including using English.
One more important thing I would like you to remember is that you are a person before you are a researcher. Please use your time well not only in research but for other things, too. This will lead to better research.
I like to watch baseball and soccer myself. There’s a special place in my heart for team play even in research, and I always get the most joy when I achieve something as a team. This is because I get the opportunity to achieve things that go beyond my own imagination. I would like to work with members of my laboratory to form an effective team.
Interviewer: Yoshiko Fujikawa, Science writer
Photographer: Mitsuhiko Kurusu, NIG ORD
July 2021
